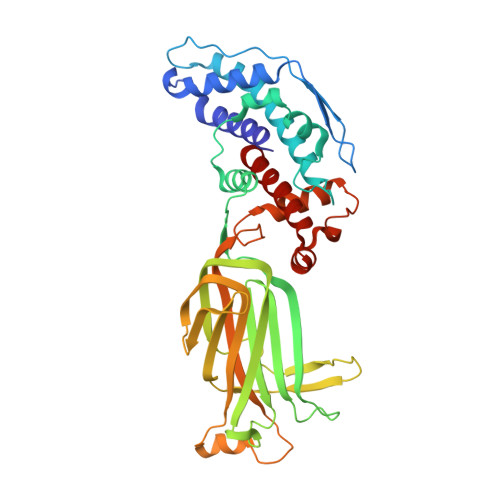
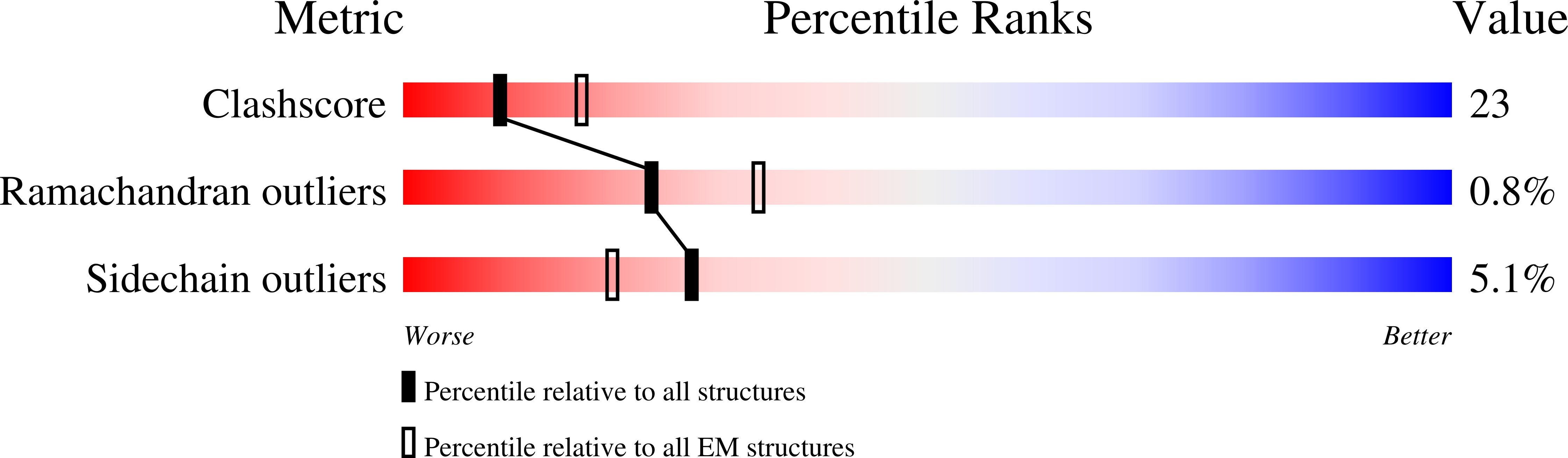Measuring the optimal exposure for single particle cryo-EM using a 2.6 angstrom reconstruction of rotavirus VP6.
Grant, T., Grigorieff, N.(2015) Elife 4: e06980-e06980
- PubMed: 26023829
- DOI: https://doi.org/10.7554/eLife.06980
- Primary Citation of Related Structures:
3J9S - PubMed Abstract:
Biological specimens suffer radiation damage when imaged in an electron microscope, ultimately limiting the attainable resolution. At a given resolution, an optimal exposure can be defined that maximizes the signal-to-noise ratio in the image. Using a 2.6 Å resolution single particle cryo-EM reconstruction of rotavirus VP6, determined from movies recorded with a total exposure of 100 electrons/Å(2), we obtained accurate measurements of optimal exposure values over a wide range of resolutions. At low and intermediate resolutions, our measured values are considerably higher than obtained previously for crystalline specimens, indicating that both images and movies should be collected with higher exposures than are generally used. We demonstrate a method of using our optimal exposure values to filter movie frames, yielding images with improved contrast that lead to higher resolution reconstructions. This 'high-exposure' technique should benefit cryo-EM work on all types of samples, especially those of relatively low-molecular mass.
Organizational Affiliation:
Janelia Research Campus, Howard Hughes Medical Institute, Ashburn, United States.