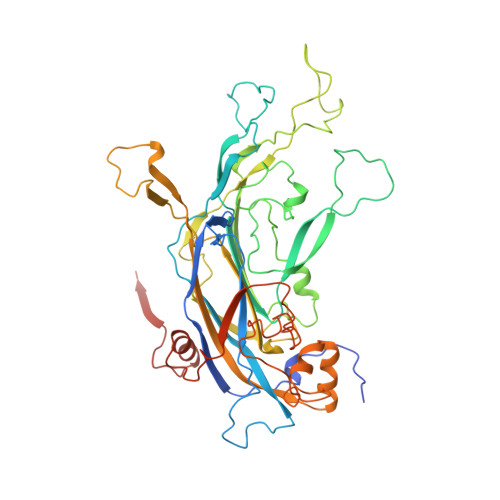
Subunit interactions in bovine papillomavirus.
Wolf, M., Garcea, R.L., Grigorieff, N., Harrison, S.C.(2010) Proc Natl Acad Sci U S A 107: 6298-6303
- PubMed: 20308582
- DOI: https://doi.org/10.1073/pnas.0914604107
- Primary Citation of Related Structures:
3IYJ - PubMed Abstract:
Papillomaviruses, members of a group of dsDNA viruses associated with epithelial growths and tumors, have compact capsids assembled from 72 pentamers of the protein L1. We have determined the structure of bovine papillomavirus by electron cryomicrosopy (cryoEM), at approximately 3.6 A resolution. The density map, obtained from single-particle analysis of approximately 4,000 particle images, shows the trace of the L1 polypeptide chain and reveals how the N- and C-terminal "arms" of a subunit (extensions from its beta-jelly-roll core) associate with a neighboring pentamer. Critical contacts come from the C-terminal arm, which loops out from the core of the subunit, forms contacts (including a disulfide) with two subunits in a neighboring pentamer, and reinserts into the pentamer from which it emanates. This trace corrects one feature of an earlier model. We discuss implications of the structure for virion assembly and for pathways of infectious viral entry. We suggest that it should be possible to obtain image reconstructions of comparable resolution from cryoEM images of asymmetric particles. From the work on papillomavirus described here, we estimate that such a reconstruction will require about 1.5 million images to achieve the same number of averaged asymmetric units; structural variability will increase this number substantially.
Organizational Affiliation:
Jack and Eileen Connors Structural Biology Laboratory, Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, 250 Longwood Avenue, Boston, MA 02115, USA.