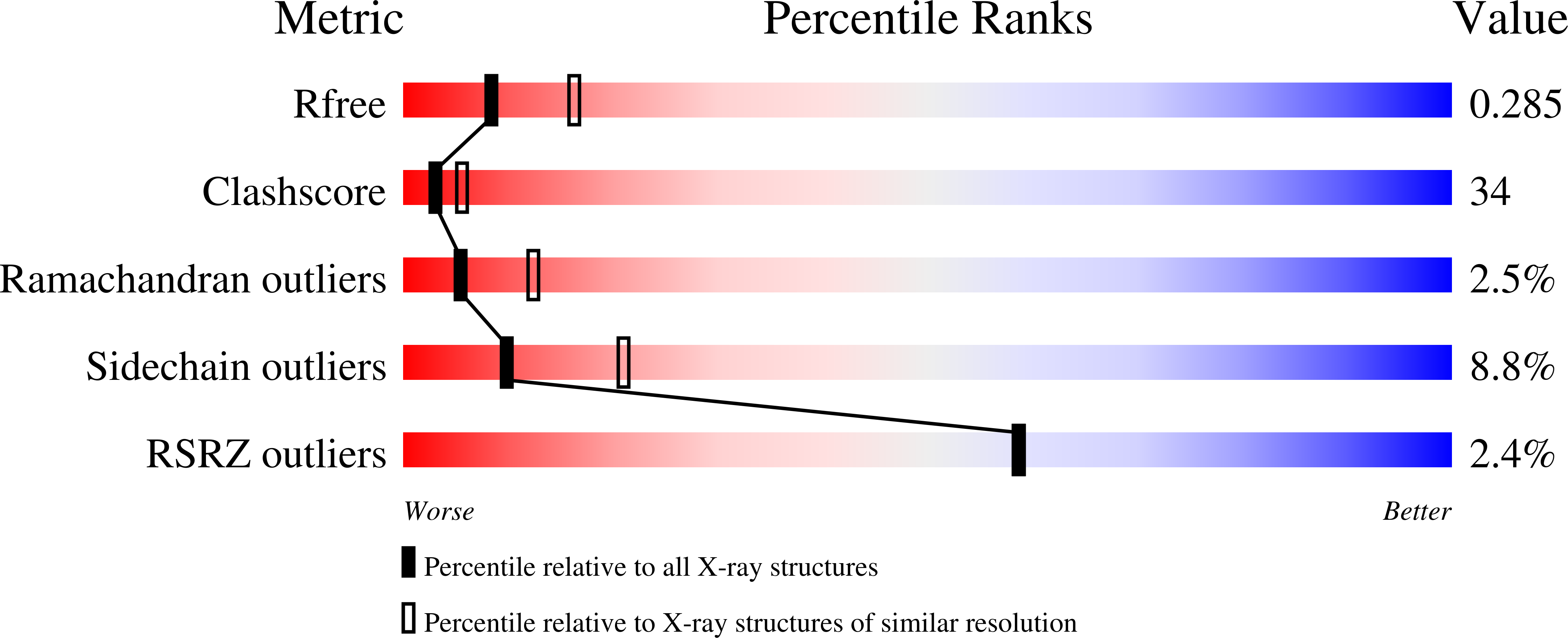
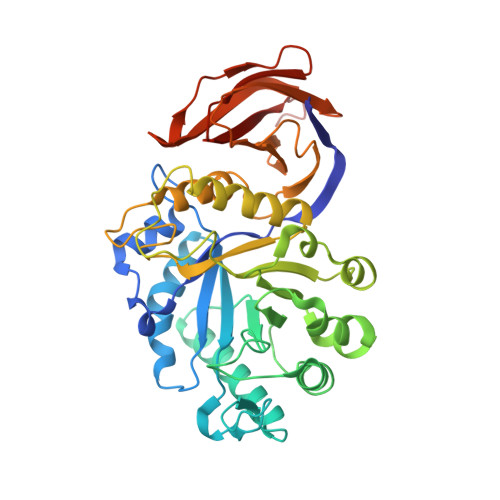
Crystallization and crystallographic analysis of Bacillus subtilis xylanase C.
St John, F.J., Godwin, D.K., Preston, J.F., Pozharski, E., Hurlbert, J.C.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 499-503
- PubMed: 19407387
- DOI: https://doi.org/10.1107/S1744309109013098
- Primary Citation of Related Structures:
3GTN - PubMed Abstract:
The recent biochemical characterization of the xylanases of glycosyl hydrolase family 5 (GH 5) has identified a distinctive endo mode of action, hydrolyzing the beta-1,4 xylan chain at a specific site directed by the position of an alpha-1,2-linked glucuronate moiety. Xylanase C (XynC), the GH 5 xylanase from Bacillus subtilis 168, has been cloned, overexpressed and crystallized. Initial data collection was performed and a preliminary model has been built into a low-quality 2.7 A resolution density map. The crystals belonged to the primitive monoclinic space group P2(1). Further screening identified an additive that resulted in large reproducible crystals. This larger more robust crystal form belonged to space group P2(1)2(1)2 and a resulting data set has been processed to 1.64 A resolution. This will be the second structure to be solved from this unique xylanase family and the first from a Gram-positive bacterium. This work may help to identify the structural determinants that allow the exceptional specificity of this enzyme and the role it plays in the biological depolymerization and processing of glucuronoxylan.
Organizational Affiliation:
University of Maryland School of Pharmacy, Department of Pharmaceutical Sciences, Baltimore, 21201, USA. fjstjohn@gmail.com