Structural insight into the inhibition of human kynurenine aminotransferase I/glutamine transaminase K
Han, Q., Robinson, H., Cai, T., Tagle, D.A., Li, J.(2009) J Med Chem 52: 2786-2793
- PubMed: 19338303
- DOI: https://doi.org/10.1021/jm9000874
- Primary Citation of Related Structures:
3FVS, 3FVU, 3FVX - PubMed Abstract:
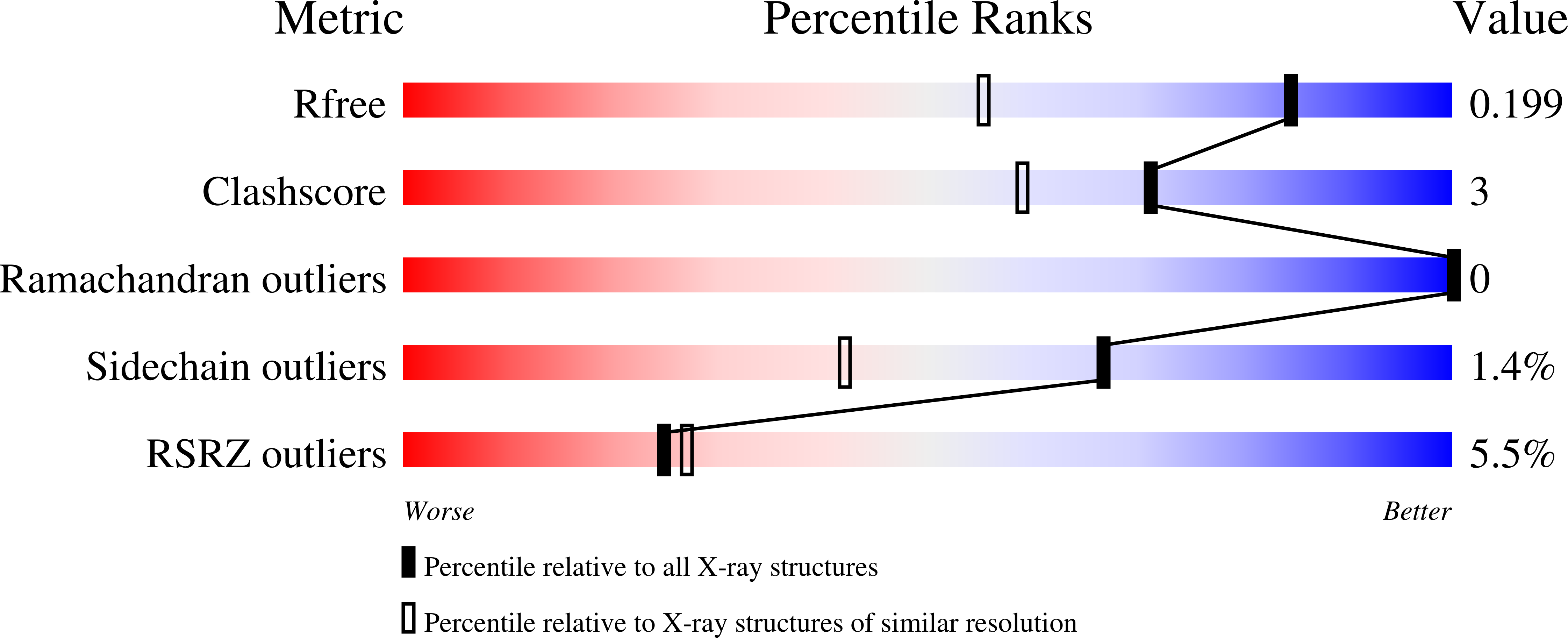
Human kynurenine aminotransferase I (hKAT I) catalyzes the formation of kynurenic acid, a neuroactive compound. Here, we report three high-resolution crystal structures (1.50-1.55 A) of hKAT I that are in complex with glycerol and each of two inhibitors of hKAT I: indole-3-acetic acid (IAC) and Tris. Because Tris is able to occupy the substrate binding position, we speculate that this may be the basis for hKAT I inhibition. Furthermore, the hKAT/IAC complex structure reveals that the binding moieties of the inhibitor are its indole ring and a carboxyl group. Six chemicals with both binding moieties were tested for their ability to inhibit hKAT I activity; 3-indolepropionic acid and DL-indole-3-lactic acid demonstrated the highest level of inhibition, and as they cannot be considered as substrates of the enzyme, these two inhibitors are promising candidates for future study. Perhaps even more significantly, we report the discovery of two different ligands located simultaneously in the hKAT I active center for the first time.
Organizational Affiliation:
Department of Biochemistry, Virginia Tech, Blacksburg, Virginia 24061, USA. qianhan@vt.edu