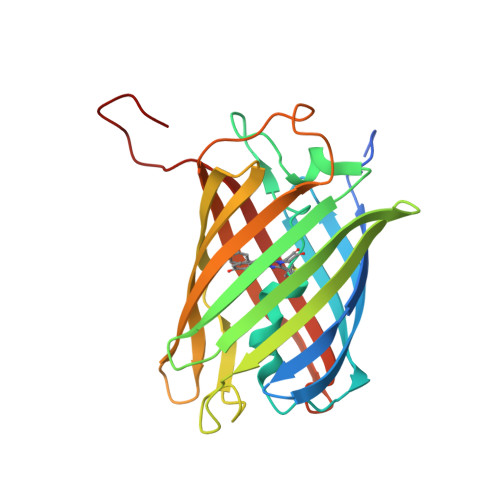
Trans-cis isomerization is responsible for the red-shifted fluorescence in variants of the red fluorescent protein eqFP611.
Nienhaus, K., Nar, H., Heilker, R., Wiedenmann, J., Nienhaus, G.U.(2008) J Am Chem Soc 130: 12578-12579
- PubMed: 18761441
- DOI: https://doi.org/10.1021/ja8046443
- Primary Citation of Related Structures:
3E5T, 3E5V, 3E5W - PubMed Abstract:
An important class of red fluorescent proteins (RFPs) feature a 2-iminomethyl-5-(4-hydroxybenzylidene)imidazolinone chromophore. Among these proteins, eqFP611 has the chromophore in a coplanar trans orientation, whereas the cis isomer is preferred by other RFPs such as DsRed and its variants. In the photoactivatable protein asFP595, the chromophore can even be switched from the nonfluorescent trans to the fluorescent cis state by light. By using X-ray crystallography, we have determined the structure of dimeric eqFP611 at high resolution (up to 1.1 A). In the far-red emitting eqFP611 variant d2RFP630, which carries an additional Asn143Ser mutation, the chromophore resides predominantly (approximately 80%) in the cis isomeric state, and in RFP639, which has Asn143Ser and Ser158Cys mutations, the chromophore is found completely in the cis form. The pronounced red shift of excitation and emission maxima of RFP639 can thus unambiguously be assigned to trans-cis isomerization of the chromophore. Among RFPs, eqFP611 is thus unique because its chromophore is highly fluorescent in both the cis and trans isomeric forms.
Organizational Affiliation:
Institute of Biophysics, University of Ulm, Albert-Einstein-Allee 11, 89081 Ulm, Germany.