Molecular basis of recognition of human osteopontin by 23C3, a potential therapeutic antibody for treatment of rheumatoid arthritis
Du, J., Hou, S., Zhong, C., Lai, Z., Yang, H., Dai, J., Zhang, D., Wang, H., Guo, Y., Ding, J.(2008) J Mol Biol 382: 835-842
- PubMed: 18694758
- DOI: https://doi.org/10.1016/j.jmb.2008.07.075
- Primary Citation of Related Structures:
3CXD, 3DSF - PubMed Abstract:
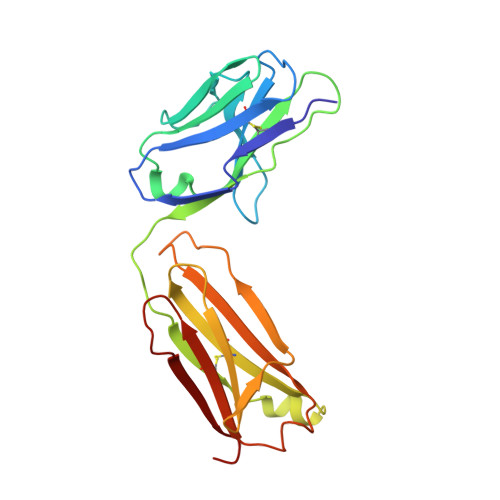
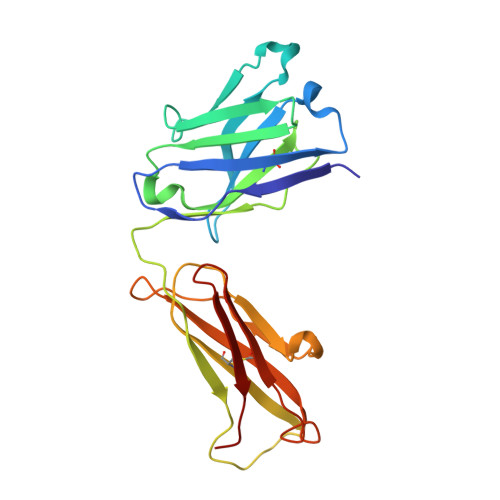
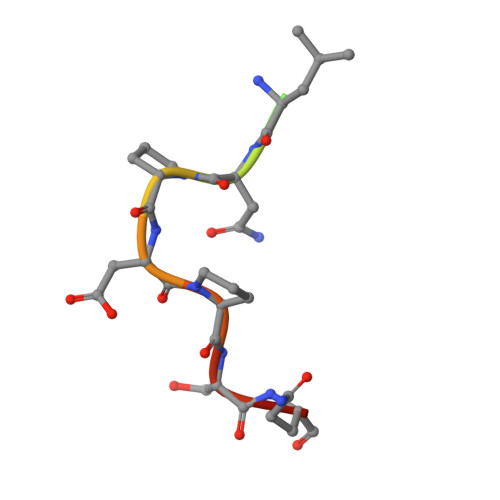
Osteopontin plays an important role in the development and perpetuation of rheumatoid arthritis (RA). Antibodies targeting osteopontin have shown promising therapeutic benefits against this disease. We have previously reported a novel anti-RA monoclonal antibody, namely, 23C3, and shown it capable of alleviating the symptoms of RA in a murine collagen-induced arthritis model, restoring the cytokine production profile in joint tissues, and reducing T-cell recall responses to collagen type II. We describe here the crystal structure of 23C3 in complex with its epitope peptide. Analyses of the complex structure reveal the molecular mechanism of osteopontin recognition by 23C3. The peptide folds into two tandem beta-turns, and two key residues of the peptide are identified to be critical for the recognition by 23C3: TrpP43 is deeply embedded into a hydrophobic pocket formed by AlaL34, TyrL36, LeuL46, TyrL49, PheL91, and MetH102 and therefore has extensive hydrophobic interactions with 23C3, while AspP47 has a network of hydrophilic interactions with residues ArgH50, ArgH52, SerH53, and AsnH56 of the antibody. Besides the complementarity-determining region loops, the framework region L2 of 23C3 is also shown to interact with the epitope peptide, which is not common in the antibody-antigen interactions and thus could be exploited in the engineering of 23C3. These results not only provide valuable information for further improvement of 23C3 such as chimerization or humanization for its therapeutic application, but also reveal the features of this specific epitope of osteopontin that may be useful for the development of new antibody drugs against RA.
Organizational Affiliation:
State Key Laboratory of Molecular Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China.