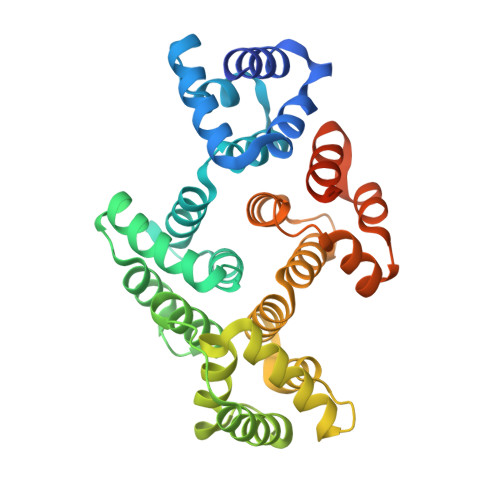
Apo and calcium-bound crystal structures of cytoskeletal protein alpha-14 giardin (annexin E1) from the intestinal protozoan parasite Giardia lamblia
Pathuri, P., Nguyen, E.T., Ozorowski, G., Svard, S.G., Luecke, H.(2009) J Mol Biol 385: 1098-1112
- PubMed: 19046974
- DOI: https://doi.org/10.1016/j.jmb.2008.11.012
- Primary Citation of Related Structures:
3CHJ, 3CHK, 3CHL - PubMed Abstract:
Alpha-14 giardin (annexin E1), a member of the alpha giardin family of annexins, has been shown to localize to the flagella of the intestinal protozoan parasite Giardia lamblia. Alpha giardins show a common ancestry with the annexins, a family of proteins most of which bind to phospholipids and cellular membranes in a Ca(2+)-dependent manner and are implicated in numerous membrane-related processes including cytoskeletal rearrangements and membrane organization. It has been proposed that alpha-14 giardin may play a significant role during the cytoskeletal rearrangement during differentiation of Giardia. To gain a better understanding of alpha-14 giardin's mode of action and its biological role, we have determined the three-dimensional structure of alpha-14 giardin and its phospholipid-binding properties. Here, we report the apo crystal structure of alpha-14 giardin determined in two different crystal forms as well as the Ca(2+)-bound crystal structure of alpha-14 giardin, refined to 1.9, 1.6 and 1.65 A, respectively. Although the overall fold of alpha-14 giardin is similar to that of alpha-11 giardin, multiwavelength anomalous dispersion phasing was required to solve the alpha-14 giardin structure, indicating significant structural differences between these two members of the alpha giardin family. Unlike most annexin structures, which typically possess N-terminal domains, alpha-14 giardin is composed of only a core domain, followed by a C-terminal extension that may serve as a ligand for binding to cytoskeletal protein partners in Giardia. In the Ca(2+)-bound structure we detected five bound calcium ions, one of which is a novel, highly coordinated calcium-binding site not previously observed in annexin structures. This novel high-affinity calcium-binding site is composed of seven protein donor groups, a feature rarely observed in crystal structures. In addition, phospholipid-binding assays suggest that alpha-14 giardin exhibits calcium-dependent binding to phospholipids that coordinate cytoskeletal disassembly/assembly during differentiation of the parasite.
Organizational Affiliation:
Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697, USA.