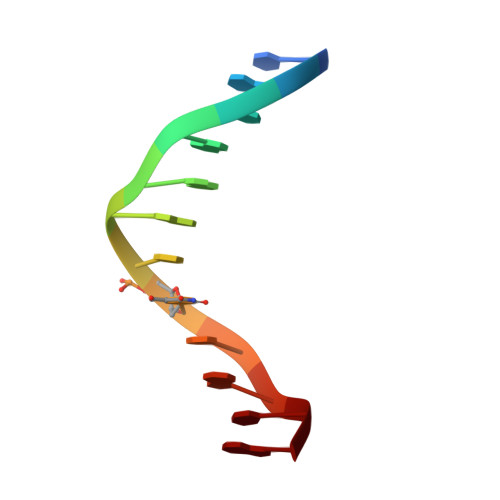
Insights into the structures of DNA damaged by hydroxyl radical: crystal structures of DNA duplexes containing 5-formyluracil
Tsunoda, M., Sakaue, T., Naito, S., Sunami, T., Abe, N., Ueno, Y., Matsuda, A., Takenaka, A.(2010) J Nucleic Acids 2010: 107289-107289
- PubMed: 20976303
- DOI: https://doi.org/10.4061/2010/107289
- Primary Citation of Related Structures:
3AJJ, 3AJK, 3AJL - PubMed Abstract:
Hydroxyl radicals are potent mutagens that attack DNA to form various base and ribose derivatives. One of the major damaged thymine derivatives is 5-formyluracil (fU), which induces pyrimidine transition during replication. In order to establish the structural basis for such mutagenesis, the crystal structures of two kinds of DNA d(CGCGRATfUCGCG) with R = A/G have been determined by X-ray crystallography. The fU residues form a Watson-Crick-type pair with A and two types of pairs (wobble and reversed wobble) with G, the latter being a new type of base pair between ionized thymine base and guanine base. In silico structural modeling suggests that the DNA polymerase can accept the reversed wobble pair with G, as well as the Watson-Crick pair with A.
Organizational Affiliation:
Faculty of Pharmacy, Iwaki Meisei University, Chuodai-Iino, Iwaki 970-8551, Japan.