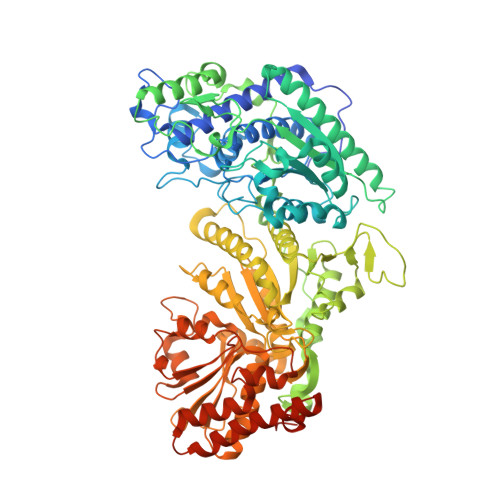
Crystal Structures of phosphoketolase: thiamine diphosphate-dependent dehydration mechanism
Suzuki, R., Katayama, T., Kim, B.-J., Wakagi, T., Shoun, H., Ashida, H., Yamamoto, K., Fushinobu, S.(2010) J Biol Chem 285: 34279-34287
- PubMed: 20739284
- DOI: https://doi.org/10.1074/jbc.M110.156281
- Primary Citation of Related Structures:
3AHC, 3AHD, 3AHE, 3AHF, 3AHG, 3AHH, 3AHI, 3AHJ - PubMed Abstract:
Thiamine diphosphate (ThDP)-dependent enzymes are ubiquitously present in all organisms and catalyze essential reactions in various metabolic pathways. ThDP-dependent phosphoketolase plays key roles in the central metabolism of heterofermentative bacteria and in the pentose catabolism of various microbes. In particular, bifidobacteria, representatives of beneficial commensal bacteria, have an effective glycolytic pathway called bifid shunt in which 2.5 mol of ATP are produced per glucose. Phosphoketolase catalyzes two steps in the bifid shunt because of its dual-substrate specificity; they are phosphorolytic cleavage of fructose 6-phosphate or xylulose 5-phosphate to produce aldose phosphate, acetyl phosphate, and H(2)O. The phosphoketolase reaction is different from other well studied ThDP-dependent enzymes because it involves a dehydration step. Although phosphoketolase was discovered more than 50 years ago, its three-dimensional structure remains unclear. In this study we report the crystal structures of xylulose 5-phosphate/fructose 6-phosphate phosphoketolase from Bifidobacterium breve. The structures of the two intermediates before and after dehydration (α,β-dihydroxyethyl ThDP and 2-acetyl-ThDP) and complex with inorganic phosphate give an insight into the mechanism of each step of the enzymatic reaction.
Organizational Affiliation:
Department of Biotechnology, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.