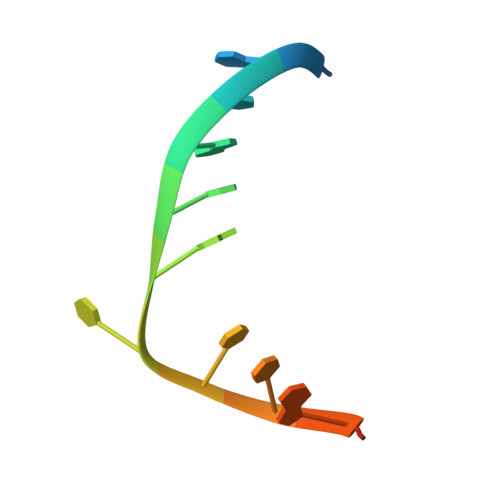
Searching for DNA lesions: structural evidence for lower- and higher-affinity DNA binding conformations of human alkyladenine DNA glycosylase.
Setser, J.W., Lingaraju, G.M., Davis, C.A., Samson, L.D., Drennan, C.L.(2012) Biochemistry 51: 382-390
- PubMed: 22148158
- DOI: https://doi.org/10.1021/bi201484k
- Primary Citation of Related Structures:
3UBY - PubMed Abstract:
To efficiently repair DNA, human alkyladenine DNA glycosylase (AAG) must search the million-fold excess of unmodified DNA bases to find a handful of DNA lesions. Such a search can be facilitated by the ability of glycosylases, like AAG, to interact with DNA using two affinities: a lower-affinity interaction in a searching process and a higher-affinity interaction for catalytic repair. Here, we present crystal structures of AAG trapped in two DNA-bound states. The lower-affinity depiction allows us to investigate, for the first time, the conformation of this protein in the absence of a tightly bound DNA adduct. We find that active site residues of AAG involved in binding lesion bases are in a disordered state. Furthermore, two loops that contribute significantly to the positive electrostatic surface of AAG are disordered. Additionally, a higher-affinity state of AAG captured here provides a fortuitous snapshot of how this enzyme interacts with a DNA adduct that resembles a one-base loop.
Organizational Affiliation:
Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, United States.