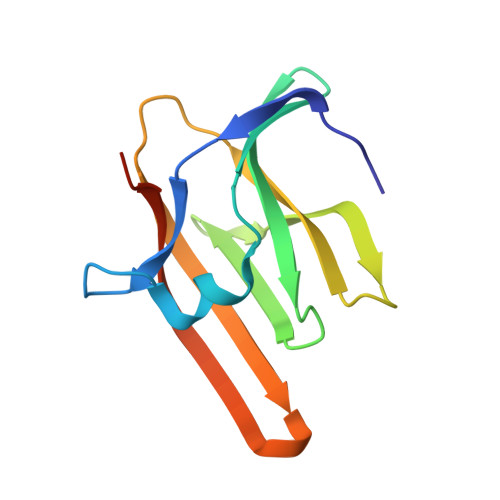
Unique Heme-Iron Coordination by the Hemoglobin Receptor IsdB of Staphylococcus aureus.
Gaudin, C.F., Grigg, J.C., Arrieta, A.L., Murphy, M.E.(2011) Biochemistry 50: 5443-5452
- PubMed: 21574663
- DOI: https://doi.org/10.1021/bi200369p
- Primary Citation of Related Structures:
3RTL, 3RUR - PubMed Abstract:
Iron is an essential requirement for life for nearly all organisms. The human pathogen Staphylococcus aureus is able to acquire iron from the heme cofactor of hemoglobin (Hb) released from lysed erythrocytes. IsdB, the predominant Hb receptor of S. aureus, is a cell wall-anchored protein that is composed of two NEAT domains. The N-terminal NEAT domain (IsdB-N1) binds Hb, and the C-terminal NEAT domain (IsdB-N2) relays heme to IsdA for transport into the cell. Here we present the 1.45 Å resolution X-ray crystal structure of the IsdB-N2-heme complex. While the structure largely conforms to the eight-strand β-sandwich fold seen in other NEAT domains such as IsdA-N and uses a conserved Tyr residue to coordinate heme-iron, a Met residue is also involved in iron coordination, resulting in a novel Tyr-Met hexacoordinate heme-iron state. The kinetics of the transfer of heme from IsdB-N2 to IsdA-N can be modeled as a two-step process. The rate of transfer of heme between the isolated NEAT domains (82 s(-1)) was found to be similar to that measured for the full-length proteins. Replacing the iron coordinating Met with Leu did not abrogate high-affinity heme binding but did reduce the heme transfer rate constant by more than half. This unusual Met-Tyr heme coordination may also bestow properties on IsdB that help it to bind heme in different oxidation states or extract heme from hemoglobin.
Organizational Affiliation:
Department of Microbiology and Immunology, Life Sciences Institute, The University of British Columbia, Vancouver, BC, Canada V6T 1Z3.