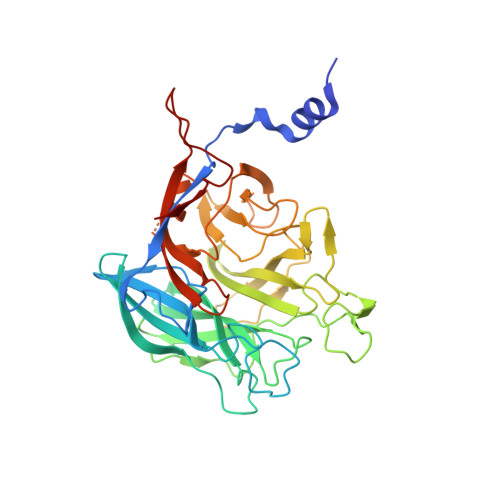
Crystal structure of a key enzyme in the agarolytic pathway, alpha-neoagarobiose hydrolase from Saccharophagus degradans 2-40
Ha, S.C., Lee, S., Lee, J., Kim, H.T., Ko, H.J., Kim, K.H., Choi, I.G.(2011) Biochem Biophys Res Commun 412: 238-244
- PubMed: 21810409
- DOI: https://doi.org/10.1016/j.bbrc.2011.07.073
- Primary Citation of Related Structures:
3R4Y, 3R4Z - PubMed Abstract:
In agarolytic microorganisms, α-neoagarobiose hydrolase (NABH) is an essential enzyme to metabolize agar because it converts α-neoagarobiose (O-3,6-anhydro-alpha-l-galactopyranosyl-(1,3)-d-galactose) into fermentable monosaccharides (d-galactose and 3,6-anhydro-l-galactose) in the agarolytic pathway. NABH can be divided into two biological classes by its cellular location. Here, we describe a structure and function of cytosolic NABH from Saccharophagus degradans 2-40 in a native protein and d-galactose complex determined at 2.0 and 1.55 Å, respectively. The overall fold is organized in an N-terminal helical extension and a C-terminal five-bladed β-propeller catalytic domain. The structure of the enzyme-ligand (d-galactose) complex predicts a +1 subsite in the substrate binding pocket. The structural features may provide insights for the evolution and classification of NABH in agarolytic pathways.
Organizational Affiliation:
Physical Bioscience Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA.