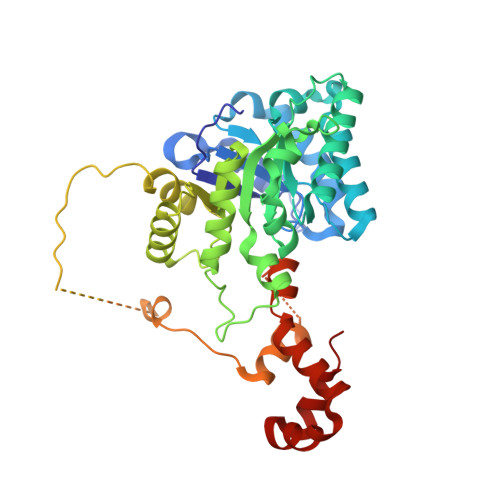
Crystal structures of a halophilic archaeal malate synthase from Haloferax volcanii and comparisons with isoforms A and G.
Bracken, C.D., Neighbor, A.M., Lamlenn, K.K., Thomas, G.C., Schubert, H.L., Whitby, F.G., Howard, B.R.(2011) BMC Struct Biol 11: 23-23
- PubMed: 21569248
- DOI: https://doi.org/10.1186/1472-6807-11-23
- Primary Citation of Related Structures:
3OYX, 3OYZ, 3PUG - PubMed Abstract:
Malate synthase, one of the two enzymes unique to the glyoxylate cycle, is found in all three domains of life, and is crucial to the utilization of two-carbon compounds for net biosynthetic pathways such as gluconeogenesis. In addition to the main isoforms A and G, so named because of their differential expression in E. coli grown on either acetate or glycolate respectively, a third distinct isoform has been identified. These three isoforms differ considerably in size and sequence conservation. The A isoform (MSA) comprises ~530 residues, the G isoform (MSG) is ~730 residues, and this third isoform (MSH-halophilic) is ~430 residues in length. Both isoforms A and G have been structurally characterized in detail, but no structures have been reported for the H isoform which has been found thus far only in members of the halophilic Archaea.
Organizational Affiliation:
Department of Physical Science, Southern Utah University, Cedar City, UT 84720-2470, USA.