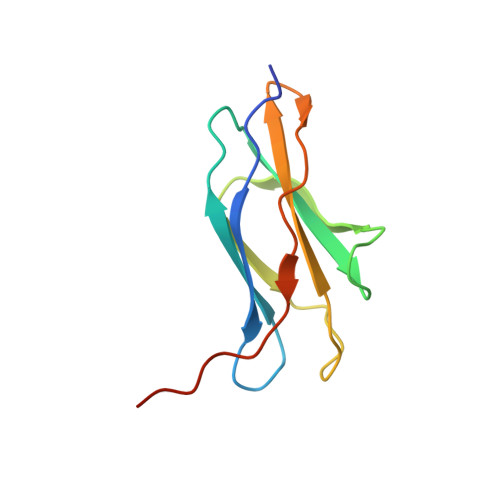
Structure of a fibronectin type III-like module from Clostridium thermocellum.
Alahuhta, M., Xu, Q., Brunecky, R., Adney, W.S., Ding, S.Y., Himmel, M.E., Lunin, V.V.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 878-880
- PubMed: 20693658
- DOI: https://doi.org/10.1107/S1744309110022529
- Primary Citation of Related Structures:
3MPC - PubMed Abstract:
The 1.6 A resolution structure of a fibronectin type III-like module from Clostridium thermocellum (PDB code 3mpc) with two molecules in the asymmetric unit is reported. The crystals used for data collection belonged to space group P2(1)2(1)2(1), with unit-cell parameters a=35.43, b=45.73, c=107.72 A, and the structure was refined to an R factor of 0.166. Structural comparisons found over 800 similar structures in the Protein Data Bank. The broad range of different proteins or protein domains with high structural similarity makes it especially demanding to classify these proteins. Previous studies of fibronectin type III-like modules have indicated that they might function as ligand-binding modules, as a compact form of peptide linkers or spacers between other domains, as cellulose-disrupting modules or as proteins that help large enzyme complexes remain soluble.
Organizational Affiliation:
BioSciences Center, National Renewable Energy Laboratory, 1617 Cole Boulevard, Golden, Colorado 80401-3305, USA.