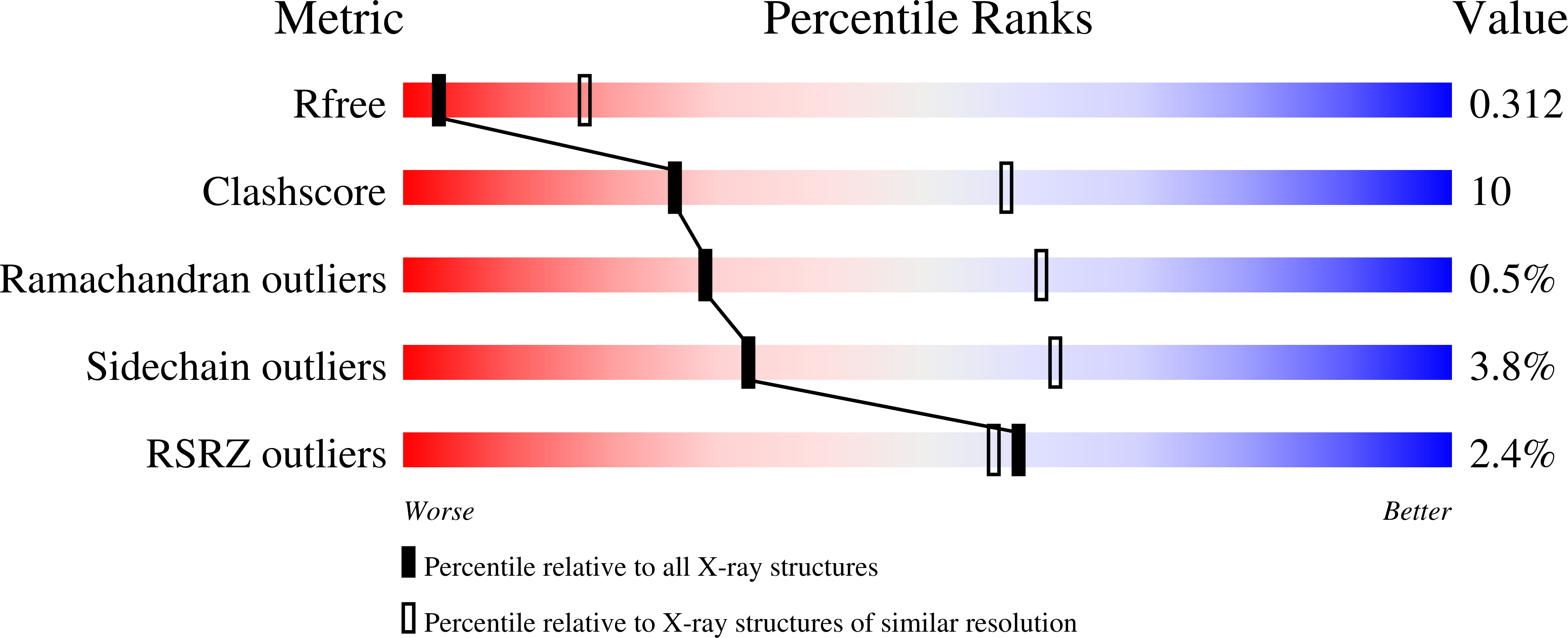
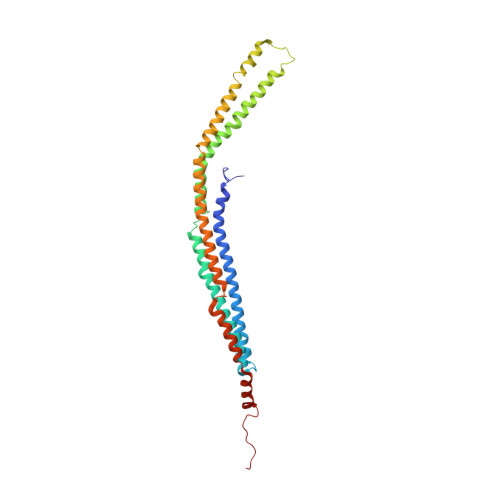
A hinge in the distal end of the PACSIN 2 F-BAR domain may contribute to membrane-curvature sensing.
Plomann, M., Wittmann, J.G., Rudolph, M.G.(2010) J Mol Biol 400: 129-136
- PubMed: 20471395
- DOI: https://doi.org/10.1016/j.jmb.2010.05.008
- Primary Citation of Related Structures:
3LLL - PubMed Abstract:
The protein kinase C and casein kinase 2 substrates in neurons (PACSINs) represent a subfamily of membrane-binding proteins characterized by an amino-terminal Bin-Amphiphysin-Rvs (F-BAR) domain. PACSINs link membrane trafficking with actin dynamics and regulate the localization of distinct cargo molecules. The F-BAR domain forms a dimer essential for lipid binding. We have obtained crystals of authentic murine PACSIN 2 that contain an ordered F-BAR domain, indicating that additional domains are flexibly connected to F-BAR. The structure shares similarity to other BAR domains and exhibits special features unique to PACSINs. These include the uneven distribution of charged residues on the concave molecular surface and a so-called wedge loop that is driven into the membrane upon binding of PACSIN. The murine PACSIN 2 F-BAR domain requires dimerization for sensing of curved membranes, and the present structure also provides a mechanism for higher-order oligomer formation. Importantly, comparison of murine with human and Drosophila PACSIN 2 F-BAR domains reveals stark differences in the orientation of distal helical segments leading to a wider crescent shape of murine PACSIN 2. We define hinge residues for these movements that may help PACSINs sense and concomitantly reinforce membrane curvature.
Organizational Affiliation:
Institute for Biochemistry II and Center for Molecular Medicine Cologne (CMMC), University of Cologne, D-50931 Cologne, Germany.