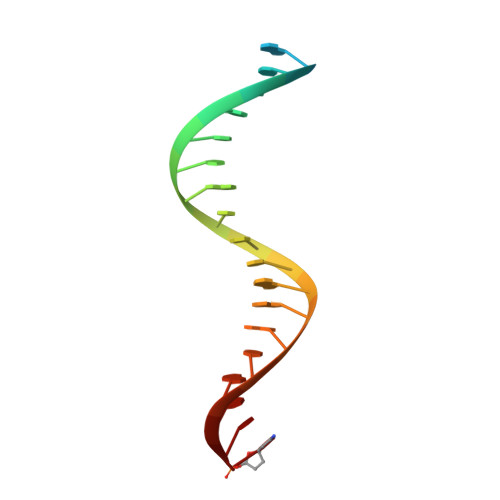
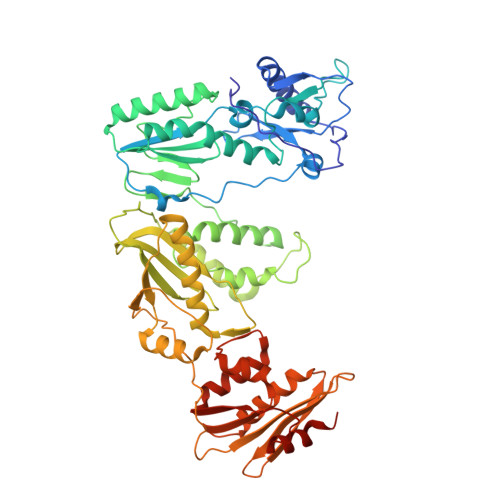
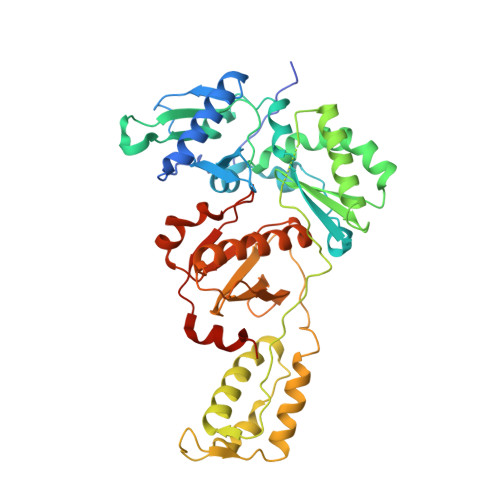
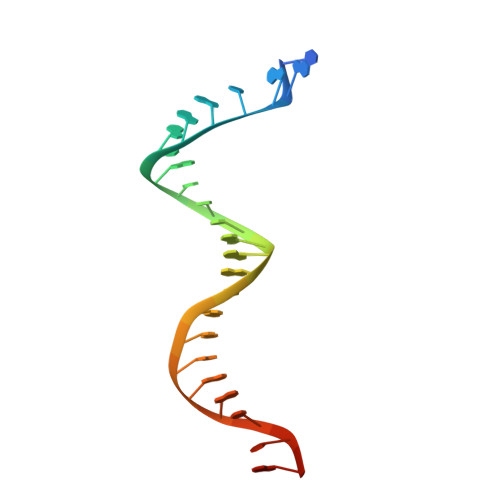
Visualizing the molecular interactions of a nucleotide analog, GS-9148, with HIV-1 reverse transcriptase-DNA complex.
Lansdon, E.B., Samuel, D., Lagpacan, L., Brendza, K.M., White, K.L., Hung, M., Liu, X., Boojamra, C.G., Mackman, R.L., Cihlar, T., Ray, A.S., McGrath, M.E., Swaminathan, S.(2010) J Mol Biol 397: 967-978
- PubMed: 20156454
- DOI: https://doi.org/10.1016/j.jmb.2010.02.019
- Primary Citation of Related Structures:
3KJV, 3KK1, 3KK2, 3KK3 - PubMed Abstract:
GS-9148 ([5-(6-amino-purin-9-yl)-4-fluoro-2,5-dihydro-furan-2-yloxymethyl]-phosphonic acid) is a dAMP (2'-deoxyadenosine monophosphate) analog that maintains its antiviral activity against drug-resistant HIV. Crystal structures for HIV-1 reverse transcriptase (RT) bound to double-stranded DNA, ternary complexes with either GS-9148-diphosphate or 2'-deoxyadenosine triphosphate (dATP), and a post-incorporation structure with GS-9148 translocated to the priming site were obtained to gain insight into the mechanism of RT inhibition. The binding of either GS-9148-diphosphate or dATP to the binary RT-DNA complex resulted in the fingers subdomain closing around the incoming substrate. This produced up to a 9 A shift in the tips of the fingers subdomain as it closed toward the palm and thumb subdomains. GS-9148-diphosphate shows a similar binding mode as dATP in the nucleotide-binding site. Residues whose mutations confer resistance to nucleotide/nucleoside RT inhibitors, such as M184, Y115, L74, and K65, show little to no shift in orientation whether GS-9148-diphosphate or dATP is bound. One difference observed in binding is the position of the central ring. The dihydrofuran ring of GS-9148-diphosphate interacts with the aromatic side chain of Y115 more than does the ribose ring of dATP, possibly picking up a favorable pi-pi interaction. The ability of GS-9148-diphosphate to mimic the active-site contacts of dATP may explain its effective inhibition of RT and maintained activity against resistance mutations. Interestingly, the 2'-fluoro moiety of GS-9148-diphosphate was found in close proximity to the Q151 side chain, potentially explaining the observed moderately reduced susceptibly to GS-9148 conferred by Q151M mutation.
Organizational Affiliation:
Gilead Sciences, Inc., 333 Lakeside Drive, Foster City, CA 94404, USA. eric.lansdon@gilead.com