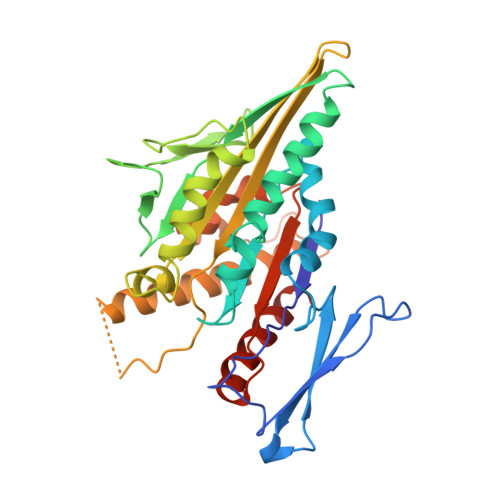
X-ray crystal structure of the yeast Kar3 motor domain complexed with Mg.ADP to 2.3 A resolution.
Gulick, A.M., Song, H., Endow, S.A., Rayment, I.(1998) Biochemistry 37: 1769-1776
- PubMed: 9485302
- DOI: https://doi.org/10.1021/bi972504o
- Primary Citation of Related Structures:
3KAR - PubMed Abstract:
The kinesin family of motor proteins, which contain a conserved motor domain of approximately 350 amino acids, generate movement against microtubules. Over 90 members of this family have been identified, including motors that move toward the minus or plus end of microtubules. The Kar3 protein from Saccharomyces cerevisiae is a minus end-directed kinesin family member that is involved in both nuclear fusion, or karyogamy, and mitosis. The Kar3 protein is 729 residues in length with the motor domain located in the C-terminal 347 residues. Recently, the three-dimensional structures of two kinesin family members have been reported. These structures include the motor domains of the plus end-directed kinesin heavy chain [Kull, F. J., et al. (1996) Nature 380, 550-555] and the minus end-directed Ncd [Sablin, E. P., et al. (1996) Nature 380, 555-559]. We now report the structure of the Kar3 protein complexed with Mg.ADP obtained from crystallographic data to 2.3 A. The structure is similar to those of the earlier kinesin family members, but shows differences as well, most notably in the length of helix alpha 4, a helix which is believed to be involved in conformational changes during the hydrolysis cycle.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison 53705, USA.