Structure-Based Design, Synthesis, and Evaluation of 2'-(2-Hydroxyethyl)-2'-deoxyadenosine and the 5'-Diphosphate Derivative as Ribonucleotide Reductase Inhibitors
Sun, D., Xu, H., Wijerathna, S.R., Dealwis, C., Lee, R.E.(2009) ChemMedChem 4: 1649-1656
- PubMed: 19681093
- DOI: https://doi.org/10.1002/cmdc.200900236
- Primary Citation of Related Structures:
3K8T - PubMed Abstract:
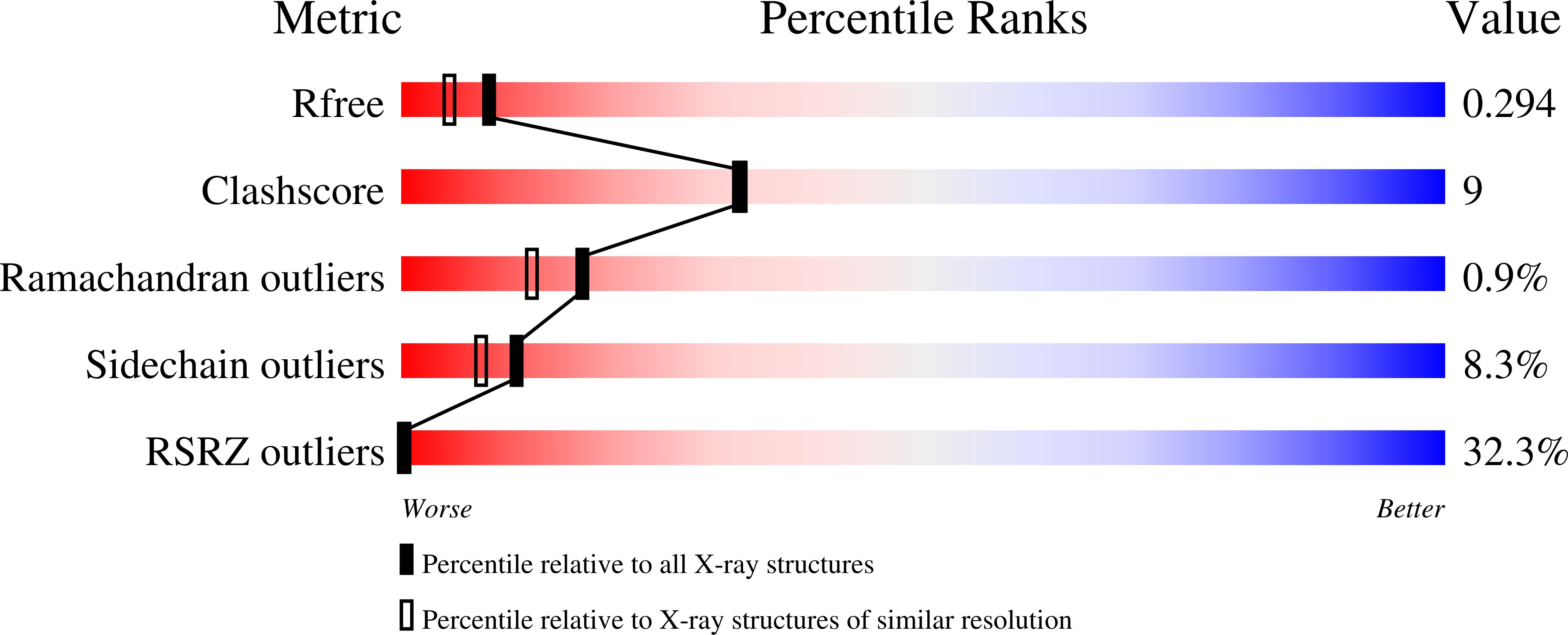
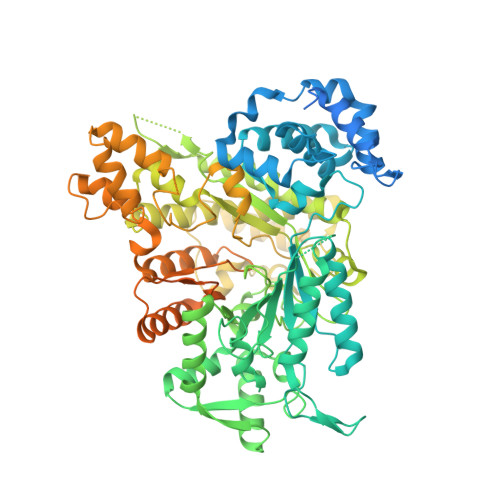
Analysis of the recently solved X-ray crystal structures of Saccharomyces cerevisiae ribonucleotide reductase I (ScRnr1) in complex with effectors and substrates led to the discovery of a conserved water molecule located at the active site that interacted with the 2'-hydroxy group of the nucleoside ribose. In this study 2'-(2-hydroxyethyl)-2'-deoxyadenosine 1 and the 5'-diphosphate derivative 2 were designed and synthesized to see if the conserved water molecule could be displaced by a hydroxymethylene group, to generate novel RNR inhibitors as potential antitumor agents. Herein we report the synthesis of analogues 1 and 2, and the co-crystal structure of adenosine diphosphate analogue 2 bound to ScRnr1, which shows the conserved water molecule is displaced as hypothesized.
Organizational Affiliation:
Department of Pharmaceutical Sciences, University of Tennessee Health Science Center, Memphis, TN 38163 (USA).