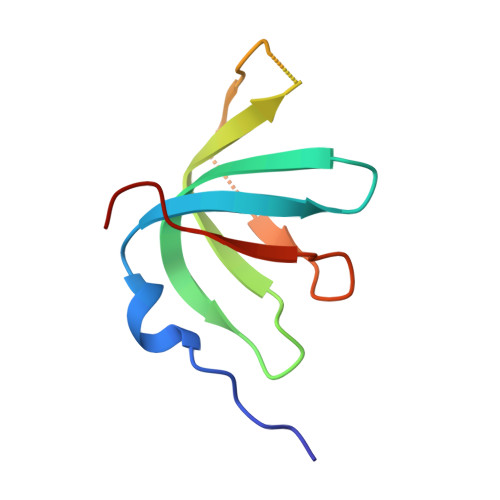
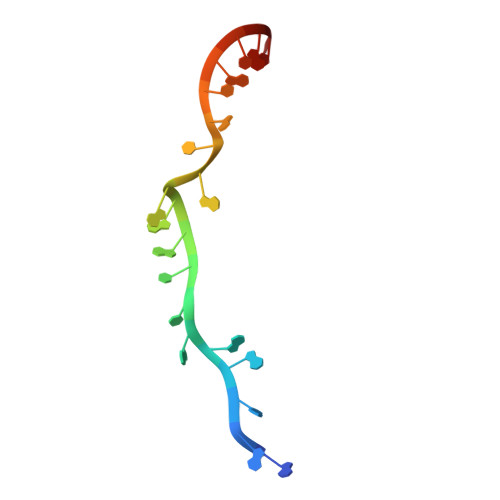
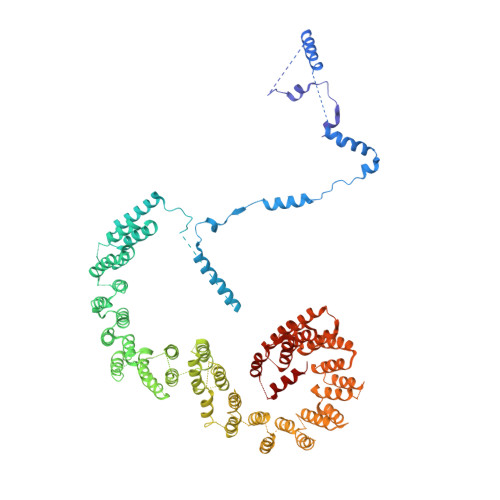
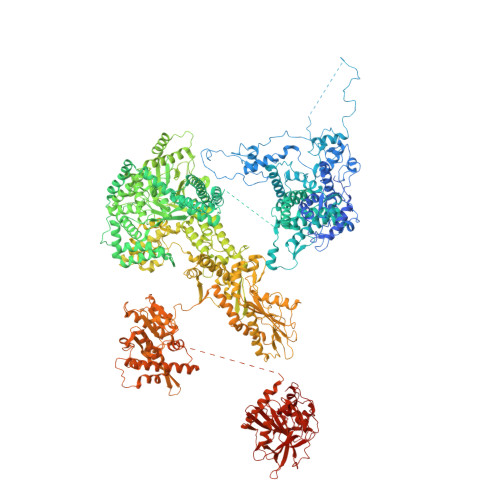
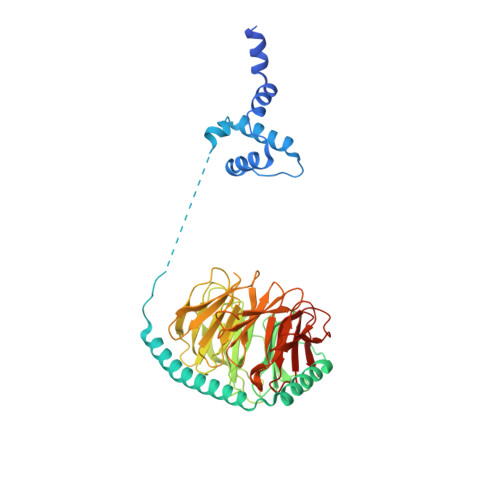
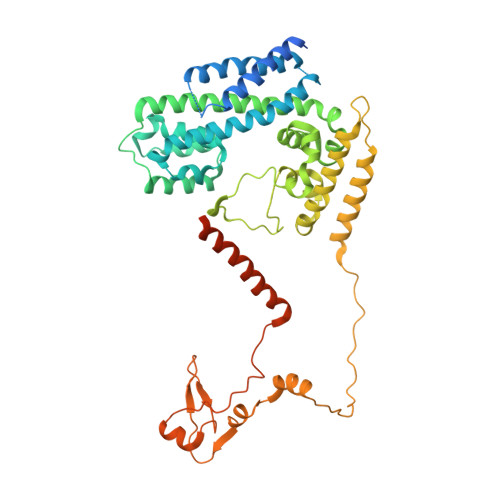
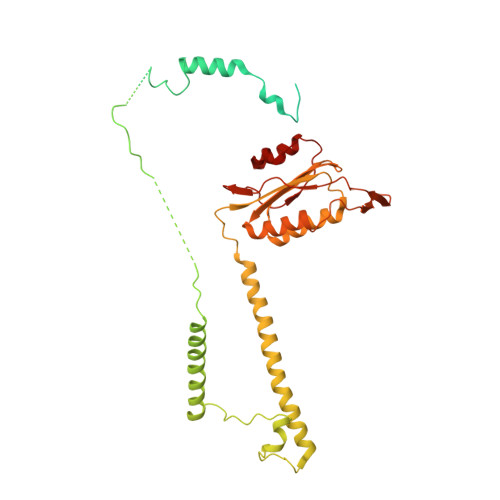
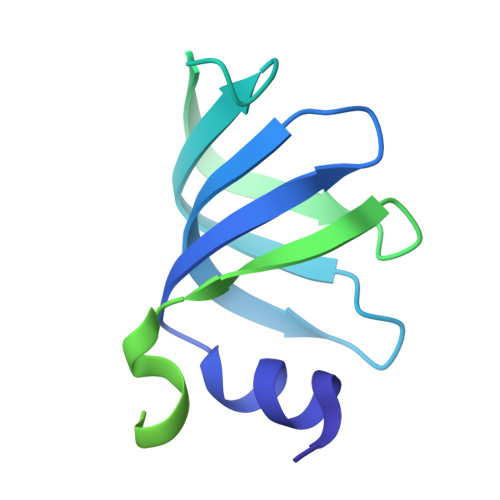
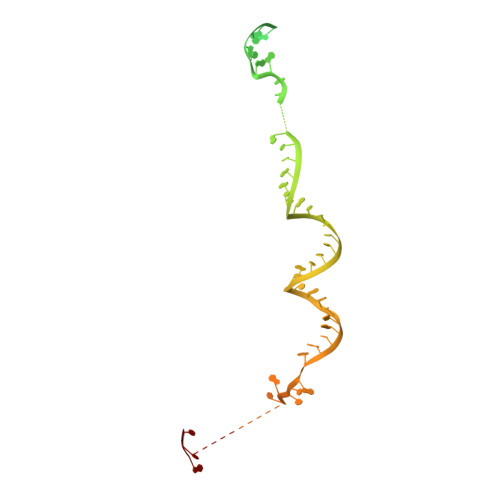
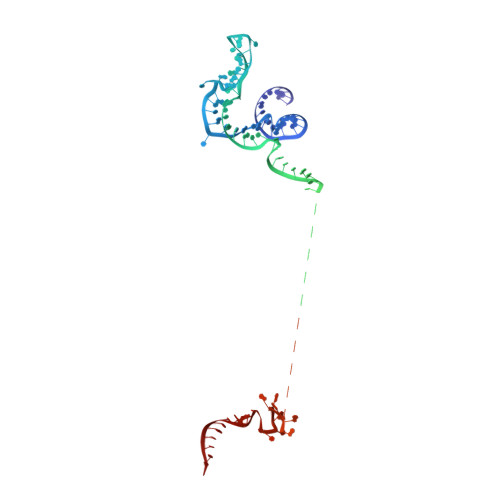
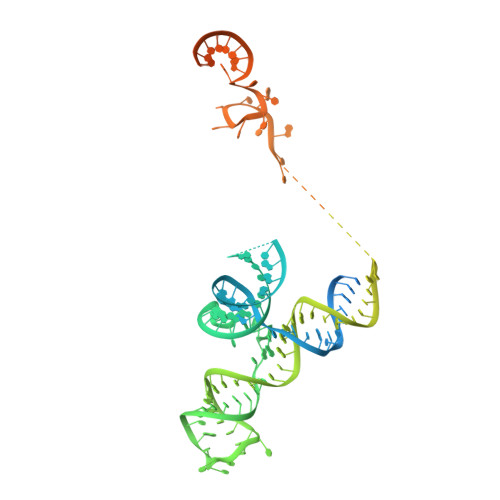
The 3.8 angstrom structure of the U4/U6.U5 tri-snRNP: Insights into spliceosome assembly and catalysis
Wan, R., Yan, C., Bai, R., Wang, L., Huang, M., Wong, C.C., Shi, Y.(2016) Science 351: 466-475
- PubMed: 26743623
- DOI: https://doi.org/10.1126/science.aad6466
- Primary Citation of Related Structures:
3JCM - PubMed Abstract:
Splicing of precursor messenger RNA is accomplished by a dynamic megacomplex known as the spliceosome. Assembly of a functional spliceosome requires a preassembled U4/U6.U5 tri-snRNP complex, which comprises the U5 small nuclear ribonucleoprotein (snRNP), the U4 and U6 small nuclear RNA (snRNA) duplex, and a number of protein factors. Here we report the three-dimensional structure of a Saccharomyces cerevisiae U4/U6.U5 tri-snRNP at an overall resolution of 3.8 angstroms by single-particle electron cryomicroscopy. The local resolution for the core regions of the tri-snRNP reaches 3.0 to 3.5 angstroms, allowing construction of a refined atomic model. Our structure contains U5 snRNA, the extensively base-paired U4/U6 snRNA, and 30 proteins including Prp8 and Snu114, which amount to 8495 amino acids and 263 nucleotides with a combined molecular mass of ~1 megadalton. The catalytic nucleotide U80 from U6 snRNA exists in an inactive conformation, stabilized by its base-pairing interactions with U4 snRNA and protected by Prp3. Pre-messenger RNA is bound in the tri-snRNP through base-pairing interactions with U6 snRNA and loop I of U5 snRNA. This structure, together with that of the spliceosome, reveals the molecular choreography of the snRNAs in the activation process of the spliceosomal ribozyme.
Organizational Affiliation:
Ministry of Education Key Laboratory of Protein Science, Tsinghua-Peking Joint Center for Life Sciences, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing 100084, China.