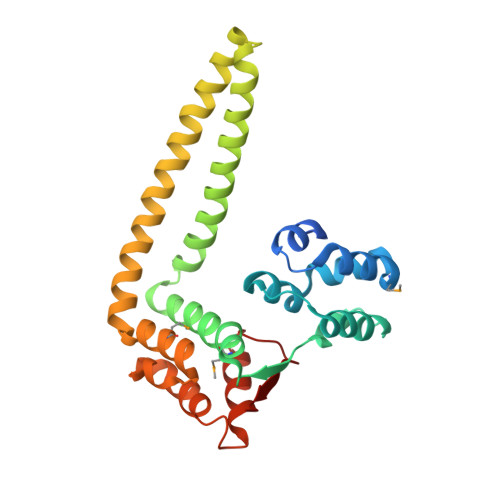
Structural evidence suggests that antiactivator ExsD from Pseudomonas aeruginosa is a DNA binding protein
Bernhards, R.C., Jing, X., Vogelaar, N.J., Robinson, H., Schubot, F.D.(2009) Protein Sci 18: 503-513
- PubMed: 19235906
- DOI: https://doi.org/10.1002/pro.48
- Primary Citation of Related Structures:
3FD9 - PubMed Abstract:
The opportunistic pathogen P. aeruginosa utilizes a type III secretion system (T3SS) to support acute infections in predisposed individuals. In this bacterium, expression of all T3SS-related genes is dependent on the AraC-type transcriptional activator ExsA. Before host contact, the T3SS is inactive and ExsA is repressed by the antiactivator protein ExsD. The repression, thought to occur through direct interactions between the two proteins, is relieved upon opening of the type III secretion (T3S) channel when secretion chaperone ExsC sequesters ExsD. We have solved the crystal structure of Delta20ExsD, a protease-resistant fragment of ExsD that lacks only the 20 amino terminal residues of the wild-type protein at 2.6 A. Surprisingly the structure revealed similarities between ExsD and the DNA binding domain of transcriptional repressor KorB. A model of an ExsD-DNA complex constructed on the basis of this homology produced a realistic complex that is supported by the prevalence of conserved residues in the putative DNA binding site and the results of differential scanning fluorimetry studies. Our findings challenge the currently held model that ExsD solely acts through interactions with ExsA and raise new questions with respect to the underlying mechanism of ExsA regulation.
Organizational Affiliation:
Department of Biological Sciences, Virginia Polytechnic Institute and State University, Blacksburg, Virginia 24060, USA.