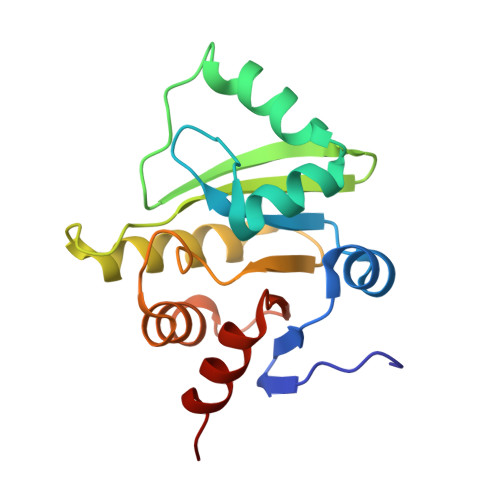
Structure of the X (ADRP) domain of nsp3 from feline coronavirus
Wojdyla, J.A., Manolaridis, I., Snijder, E.J., Gorbalenya, A.E., Coutard, B., Piotrowski, Y., Hilgenfeld, R., Tucker, P.A.(2009) Acta Crystallogr D Biol Crystallogr 65: 1292-1300
- PubMed: 19966415
- DOI: https://doi.org/10.1107/S0907444909040074
- Primary Citation of Related Structures:
3ETI, 3EW5, 3JZT - PubMed Abstract:
The structure of the X (or ADRP) domain of a pathogenic variant of feline coronavirus (FCoV) has been determined in tetragonal and cubic crystal forms to 3.1 and 2.2 A resolution, respectively. In the tetragonal crystal form, glycerol-3-phosphate was observed in the ADP-ribose-binding site. Both crystal forms contained large solvent channels and had a solvent content of higher than 70%. Only very weak binding of this domain to ADP-ribose was detected in vitro. However, the structure with ADP-ribose bound was determined in the cubic crystal form at 3.9 A resolution. The structure of the FCoV X domain had the expected macro-domain fold and is the first structure of this domain from a coronavirus belonging to subgroup 1a.
Organizational Affiliation:
EMBL Hamburg Outstation, c/o DESY, D-22603 Hamburg, Germany.