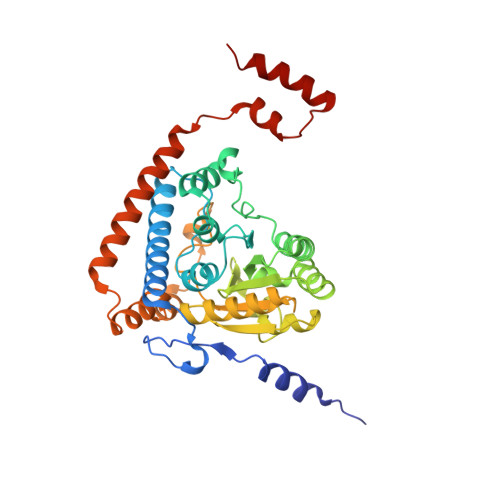
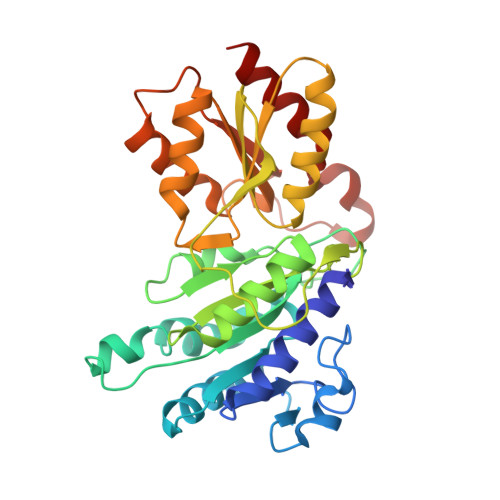
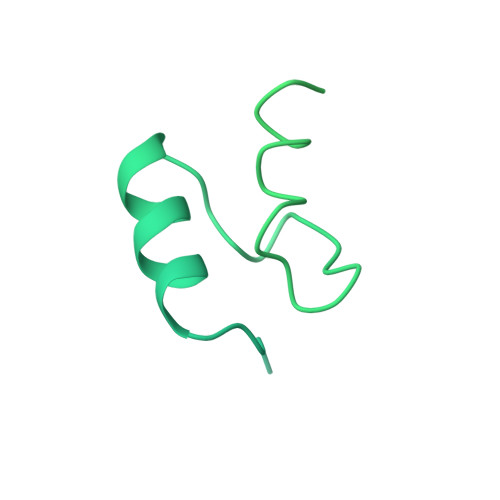
Snapshots of catalysis in the e1 subunit of the pyruvate dehydrogenase multienzyme complex
Pei, X.Y., Titman, C.M., Frank, R.A., Leeper, F.J., Luisi, B.F.(2008) Structure 16: 1860-1872
- PubMed: 19081062
- DOI: https://doi.org/10.1016/j.str.2008.10.009
- Primary Citation of Related Structures:
3DUF, 3DV0, 3DVA - PubMed Abstract:
The pyruvate dehydrogenase multienzyme assembly (PDH) generates acetyl coenzyme A and reducing equivalents from pyruvate in a multiple-step process that is a nexus of central metabolism. We report crystal structures of the Geobacillus stearothermophilus PDH E1p subunit with ligands that mimic the prereaction complex and the postdecarboxylation product. The structures implicate residues that help to orient substrates, nurture intermediates, and organize surface loops so that they can engage a mobile lipoyl domain that receives the acetyl group and shuttles it to the next active site. The structural and enzymatic data suggest that H128beta performs a dual role: first, as electrostatic catalyst of the reaction of pyruvate with the thiamine cofactor; and second, as a proton donor in the second reaction of acetyl group with the lipoate. We also identify I206alpha as a key residue in mediating the conformation of active-site loops. We propose that a simple conformational flip of the H271alpha side chain assists transfer of the acetyl group from thiamine cofactor to lipoyl domain in synchrony with reduction of the dithiolane ring.
Organizational Affiliation:
Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA, UK.