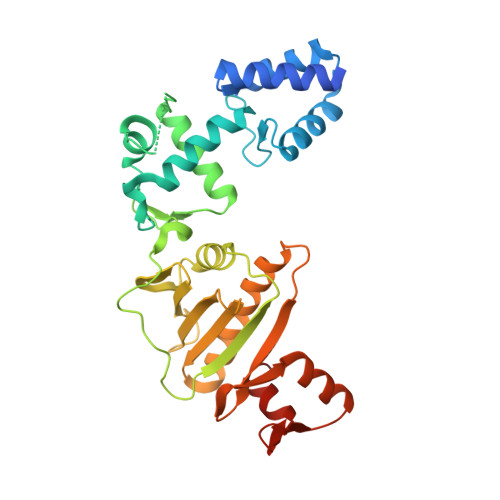
Structure of the Roc-COR domain tandem of C. tepidum, a prokaryotic homologue of the human LRRK2 Parkinson kinase
Gotthardt, K., Weyand, M., Kortholt, A., Van Haastert, P.J., Wittinghofer, A.(2008) EMBO J 27: 2239-2249
- PubMed: 18650931
- DOI: https://doi.org/10.1038/emboj.2008.150
- Primary Citation of Related Structures:
3DPT, 3DPU - PubMed Abstract:
Ras of complex proteins (Roc) belongs to the superfamily of Ras-related small G-proteins that always occurs in tandem with the C-terminal of Roc (COR) domain. This Roc-COR tandem is found in the bacterial and eukaryotic world. Its most prominent member is the leucine-rich repeat kinase LRRK2, which is mutated and activated in Parkinson patients. Here, we investigated biochemically and structurally the Roco protein from Chlorobium tepidum. We show that Roc is highly homologous to Ras, whereas the COR domain is a dimerisation device. The juxtaposition of the G-domains and mutational analysis suggest that the Roc GTPase reaction is stimulated and/or regulated by dimerisation in a nucleotide-dependent manner. The region most conserved between bacteria and man is the interface between Roc and COR, where single-point Parkinson mutations of the Roc and COR domains are in close proximity. The analogous mutations in C. tepidum Roc-COR decrease the GTPase reaction rate, most likely due to a modification of the interaction between the Roc and COR domains.
Organizational Affiliation:
Department of Structural Biology, Max-Planck-Institut for Molecular Physiology, Dortmund, Germany.