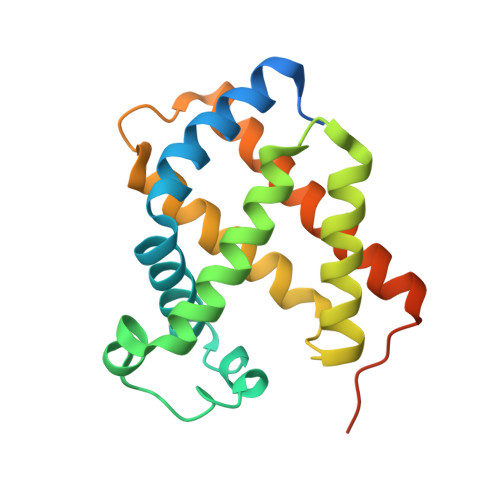
Crystal structure of the carbon monoxide complex of human cytoglobin
Makino, M., Sawai, H., Shiro, Y., Sugimoto, H.(2011) Proteins 79: 1143-1153
- PubMed: 21254233
- DOI: https://doi.org/10.1002/prot.22950
- Primary Citation of Related Structures:
3AG0 - PubMed Abstract:
Cytoglobin (Cgb) is a vertebrate heme-containing globin-protein expressed in a broad range of mammalian tissues. Unlike myoglobin, Cgb displays a hexa-coordinated (bis-hystidyl) heme iron atom, having the heme distal His81(E7) residue as the endogenous sixth ligand. In the present study, we crystallized human Cgb in the presence of a reductant Na₂S₂O₄ under a carbon monoxide (CO) atmosphere, and determined the crystal structure at 2.6 A resolution. The CO ligand occupies the sixth axial position of the heme ferrous iron. Eventually, the imidazole group of His81(E7) is expelled from the sixth position and swings out of the distal heme pocket. The flipping motion of the His81 imidazole group accompanies structural readjustments of some residues (Gln62, Phe63, Gln72, and Ser75) in both the CD-corner and D-helix regions of Cgb. On the other hand, no significant structural changes were observed in other Cgb regions, for example, on the proximal side. These structural alterations that occurred as a result of exogenous ligand (CO) binding are clearly different from those observed in other vertebrate hexa-coordinated globins (mouse neuroglobin, Drosophila melanogaster hemoglobin) and penta-coordinated sperm whale myoglobin. The present study provides the structural basis for further discussion of the unique ligand-binding properties of Cgb.
Organizational Affiliation:
RIKEN SPring-8 Center, Sayo, Hyogo 679-5148, Japan.