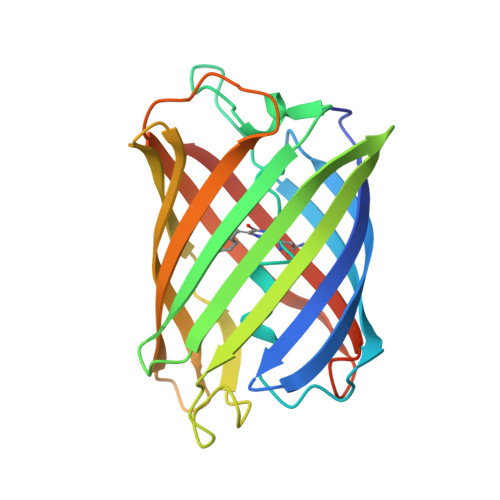
The structure of mAG, a monomeric mutant of the green fluorescent protein Azami-Green, reveals the structural basis of its stable green emission
Ebisawa, T., Yamamura, A., Kameda, Y., Hayakawa, K., Nagata, K., Tanokura, M.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 485-489
- PubMed: 20445241
- DOI: https://doi.org/10.1107/S1744309110011127
- Primary Citation of Related Structures:
3ADF - PubMed Abstract:
Monomeric Azami-Green (mAG) from the stony coral Galaxea fascicularis is the first known monomeric green-emitting fluorescent protein that is not a variant of Aequorea victoria green fluorescent protein (avGFP). These two green fluorescent proteins are only 27% identical in their amino-acid sequences. mAG is more similar in its amino-acid sequence to four fluorescent proteins: Dendra2 (a green-to-red irreversibly photoconverting fluorescent protein), Dronpa (a bright-and-dark reversibly photoswitchable fluorescent protein), KikG (a tetrameric green-emitting fluorescent protein) and Kaede (another green-to-red irreversibly photoconverting fluorescent protein). To reveal the structural basis of stable green emission by mAG, the 2.2 A crystal structure of mAG has been determined and compared with the crystal structures of avGFP, Dronpa, Dendra2, Kaede and KikG. The structural comparison revealed that the chromophore formed by Gln62-Tyr63-Gly64 (QYG) and the fixing of the conformation of the imidazole ring of His193 by hydrogen bonds and van der Waals contacts involving His193, Arg66 and Thr69 are likely to be required for the stable green emission of mAG. The crystal structure of mAG will contribute to the design and development of new monomeric fluorescent proteins with faster maturation, brighter fluorescence, improved photostability, new colours and other preferable properties as alternatives to avGFP and its variants.
Organizational Affiliation:
Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo, Japan.