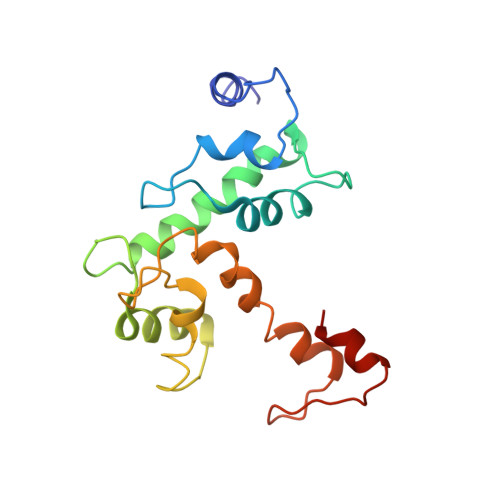
Crystallization and X-ray diffraction analysis of N-terminally truncated human ALG-2
Suzuki, H., Kawasaki, M., Kakiuchi, T., Shibata, H., Wakatsuki, S., Maki, M.(2008) Acta Crystallogr Sect F Struct Biol Cryst Commun 64: 974-977
- PubMed: 18997320
- DOI: https://doi.org/10.1107/S1744309108030297
- Primary Citation of Related Structures:
2ZRS, 2ZRT - PubMed Abstract:
ALG-2 (apoptosis-linked gene 2) is an apoptosis-linked calcium-binding protein with five EF-hand motifs in the C-terminal region. N-terminally truncated ALG-2 (des3-23ALG-2) was crystallized by the vapour-diffusion method in buffer consisting of either 50 mM MES pH 6.5, 12.5%(v/v) 2-propanol and 150 mM calcium acetate or 100 mM MES pH 6.0, 15%(v/v) ethanol and 200 mM zinc acetate. Crystals of the Ca(2+)-bound form belonged to space group P2(1)2(1)2(1), with unit-cell parameters a = 54.8, b = 154.4, c = 237.7 A, alpha = beta = gamma = 90 degrees , and diffracted to 3.1 A resolution. Crystals of the Zn(2+)-bound form belonged to space group P2(1)2(1)2(1), with unit-cell parameters a = 52.8, b = 147.5, c = 230.7 A, alpha = beta = gamma = 90 degrees , and diffracted to 3.3 A resolution. The structures of the Ca(2+)-bound form and the Zn(2+)-bound form were solved by the molecular-replacement method. Although both crystals contained eight ALG-2 molecules per asymmetric unit, the metal-ion locations and octameric arrangements were found to be significantly different.
Organizational Affiliation:
Department of Applied Molecular Biosciences, Graduate School of Bioagricultural Sciences, Nagoya University, Nagoya 464-8601, Japan.