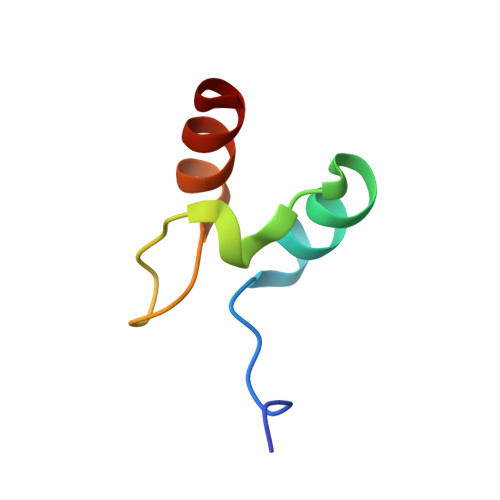
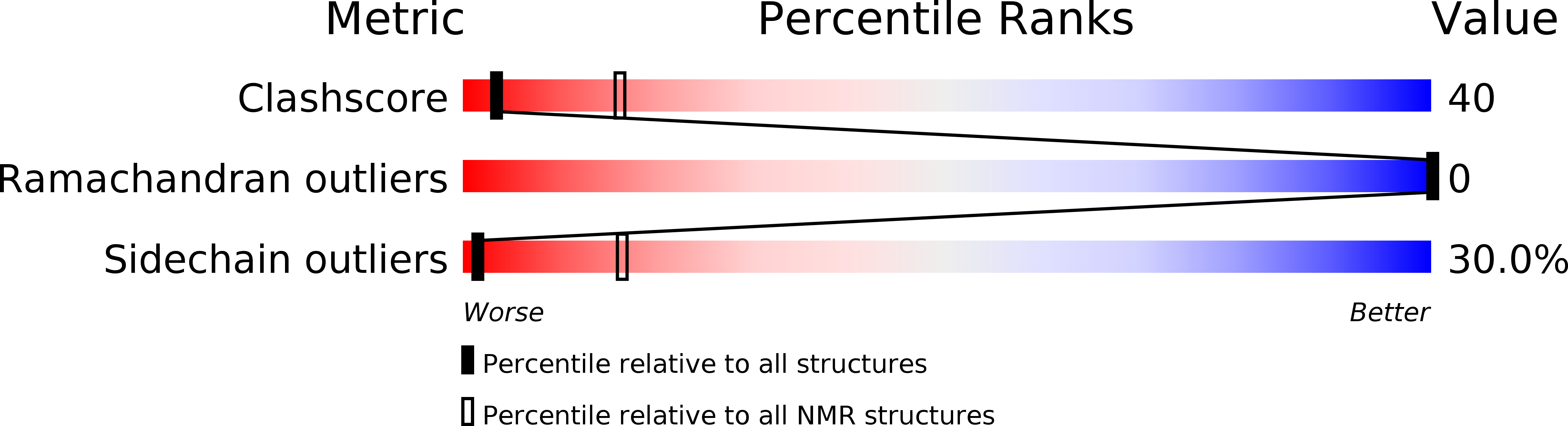The Folding Mechanism of Bbl: Plasticity of Transition-State Structure Observed within an Ultrafast Folding Protein Family.
Neuweiler, H., Sharpe, T.D., Rutherford, T.J., Johnson, C.M., Allen, M.D., Ferguson, N., Fersht, A.R.(2009) J Mol Biol 390: 1060
- PubMed: 19445954
- DOI: https://doi.org/10.1016/j.jmb.2009.05.011
- Primary Citation of Related Structures:
2WXC - PubMed Abstract:
Studies on members of protein families with similar structures but divergent sequences provide insights into the effects of sequence composition on the mechanism of folding. Members of the peripheral subunit-binding domain (PSBD) family fold ultrafast and approach the smallest size for cooperatively folding proteins. Phi-Value analysis of the PSBDs E3BD and POB reveals folding via nucleation-condensation through structurally very similar, polarized transition states. Here, we present a Phi-value analysis of the family member BBL and found that it also folds by a nucleation-condensation mechanism. The mean Phi values of BBL, E3BD, and POB were near identical, indicating similar fractions of non-covalent interactions being formed in the transition state. Despite the overall conservation of folding mechanism in this protein family, however, the pattern of Phi values determined for BBL revealed a larger dispersion of the folding nucleus across the entire structure, and the transition state was less polarized. The observed plasticity of transition-state structure can be rationalized by the different helix-forming propensities of PSBD sequences. The very strong helix propensity in the first helix of BBL, relative to E3BD and POB, appears to recruit more structure formation in that helix in the transition state at the expense of weaker interactions in the second helix. Differences in sequence composition can modulate transition-state structure of even the smallest natural protein domains.
Organizational Affiliation:
MRC Centre for Protein Engineering, Cambridge, UK.