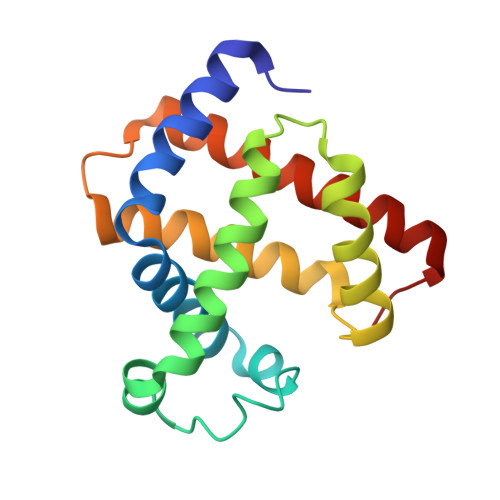
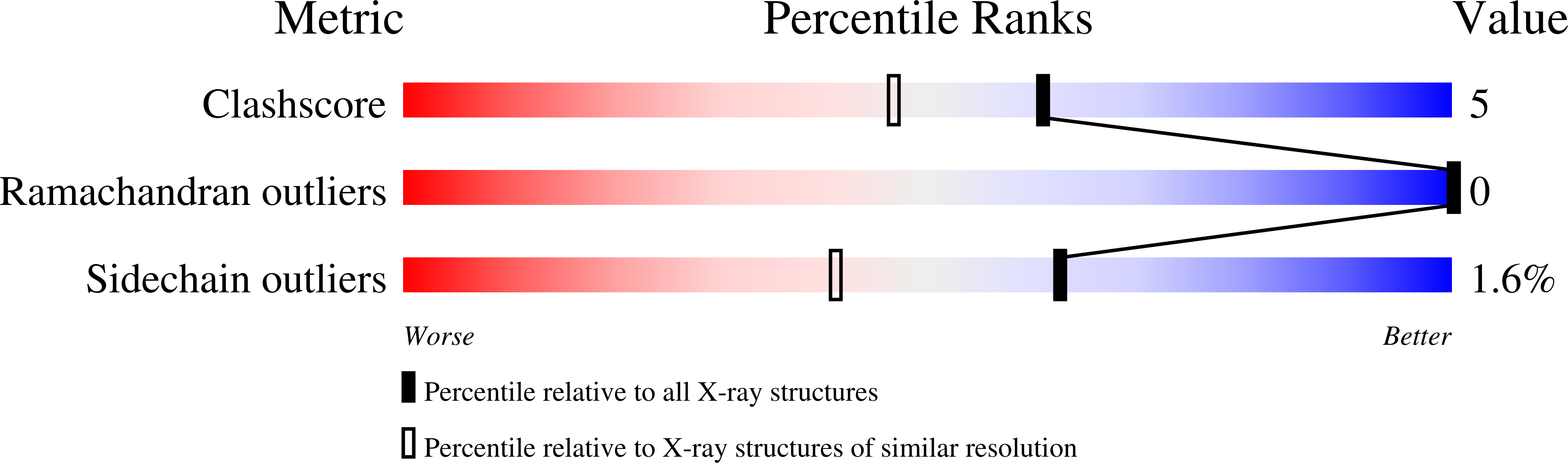The Crystal Structure of Peroxymyoglobin Generated Through Cryoradiolytic Reduction of Myoglobin Compound III During Data Collection.
Hersleth, H.-P., Hsiao, Y., Ryde, U., Gorbitz, C.H., Andersson, K.K.(2008) Biochem J 412: 257
- PubMed: 18215120
- DOI: https://doi.org/10.1042/BJ20070921
- Primary Citation of Related Structures:
2VLX, 2VLY, 2VLZ, 2VM0 - PubMed Abstract:
Myoglobin has the ability to react with hydrogen peroxide, generating high-valent complexes similar to peroxidases (compounds I and II), and in the presence of excess hydrogen peroxide a third intermediate, compound III, with an oxymyoglobin-type structure is generated from compound II. The compound III is, however, easily one-electron reduced to peroxymyoglobin by synchrotron radiation during crystallographic data collection. We have generated and solved the 1.30 A (1 A=0.1 nm) resolution crystal structure of the peroxymyoglobin intermediate, which is isoelectric to compound 0 and has a Fe-O distance of 1.8 A and O-O bond of 1.3 A in accordance with a Fe(II)-O-O- (or Fe(III)-O-O2-) structure. The generation of the peroxy intermediate through reduction of compound III by X-rays shows the importance of using single-crystal microspectrophotometry when doing crystallography on metalloproteins. After having collected crystallographic data on a peroxy-generated myoglobin crystal, we were able (by a short annealing) to break the O-O bond leading to formation of compound II. These results indicate that the cryoradiolytic-generated peroxymyoglobin is biologically relevant through its conversion into compound II upon heating. Additionally, we have observed that the Xe1 site is occupied by a water molecule, which might be the leaving group in the compound II to compound III reaction.
Organizational Affiliation:
University of Oslo, Department of Molecular Biosciences, P.O. Box 1041 Blindern, N-0316 Oslo, Norway.