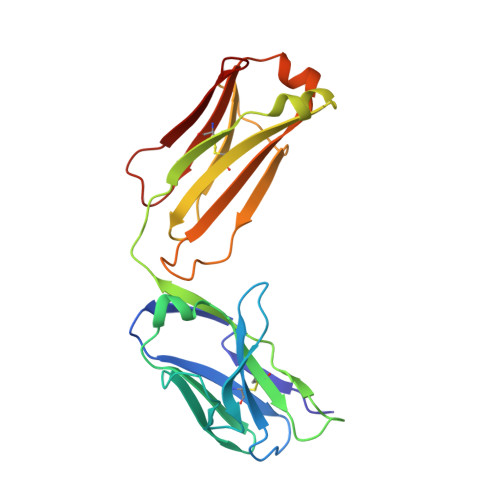
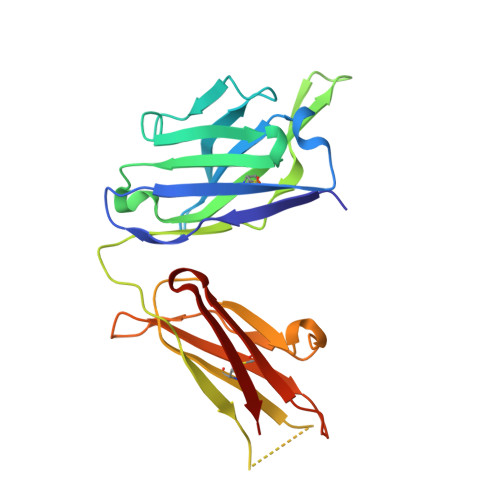
Structure of antibody F425-B4e8 in complex with a V3 peptide reveals a new binding mode for HIV-1 neutralization.
Bell, C.H., Pantophlet, R., Schiefner, A., Cavacini, L.A., Stanfield, R.L., Burton, D.R., Wilson, I.A.(2008) J Mol Biol 375: 969-978
- PubMed: 18068724
- DOI: https://doi.org/10.1016/j.jmb.2007.11.013
- Primary Citation of Related Structures:
2QSC - PubMed Abstract:
F425-B4e8 (B4e8) is a monoclonal antibody isolated from a human immunodeficiency virus type 1 (HIV-1)-infected individual that recognizes the V3 variable loop on the gp120 subunit of the viral envelope spike. B4e8 neutralizes a subset of HIV-1 primary isolates from subtypes B, C and D, which places this antibody among the very few human anti-V3 antibodies with notable cross-neutralizing activity. Here, the crystal structure of the B4e8 Fab' fragment in complex with a 24-mer V3 peptide (RP142) at 2.8 A resolution is described. The complex structure reveals that the antibody recognizes a novel V3 loop conformation, featuring a five-residue alpha-turn around the conserved GPGRA apex of the beta-hairpin loop. In agreement with previous mutagenesis analyses, the Fab' interacts primarily with V3 through side-chain contacts with just two residues, Ile(P309) and Arg(P315), while the remaining contacts are to the main chain. The structure helps explain how B4e8 can tolerate a certain degree of sequence variation within V3 and, hence, is able to neutralize an appreciable number of different HIV-1 isolates.
Organizational Affiliation:
Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.