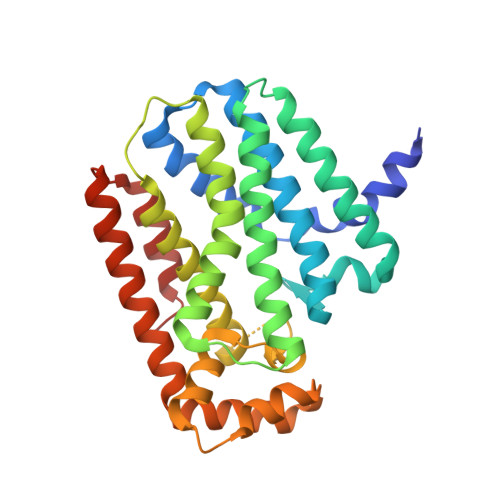
The crystal structure of human geranylgeranyl pyrophosphate synthase reveals a novel hexameric arrangement and inhibitory product binding
Kavanagh, K.L., Dunford, J.E., Bunkoczi, G., Russell, R.G., Oppermann, U.(2006) J Biol Chem 281: 22004-22012
- PubMed: 16698791
- DOI: https://doi.org/10.1074/jbc.M602603200
- Primary Citation of Related Structures:
2Q80 - PubMed Abstract:
Modification of GTPases with isoprenoid molecules derived from geranylgeranyl pyrophosphate or farnesyl pyrophosphate is an essential requisite for cellular signaling pathways. The synthesis of these isoprenoids proceeds in mammals through the mevalonate pathway, and the final steps in the synthesis are catalyzed by the related enzymes farnesyl pyrophosphate synthase and geranylgeranyl pyrophosphate synthase. Both enzymes play crucial roles in cell survival, and inhibition of farnesyl pyrophosphate synthase by nitrogen-containing bisphosphonates is an established concept in the treatment of bone disorders such as osteoporosis or certain forms of cancer in bone. Here we report the crystal structure of human geranylgeranyl pyrophosphate synthase, the first mammalian ortholog to have its x-ray structure determined. It reveals that three dimers join together to form a propeller-bladed hexameric molecule with a mass of approximately 200 kDa. Structure-based sequence alignments predict this quaternary structure to be restricted to mammalian and insect orthologs, whereas fungal, bacterial, archaeal, and plant forms exhibit the dimeric organization also observed in farnesyl pyrophosphate synthase. Geranylgeranyl pyrophosphate derived from heterologous bacterial expression is tightly bound in a cavity distinct from the chain elongation site described for farnesyl pyrophosphate synthase. The structure most likely represents an inhibitory complex, which is further corroborated by steady-state kinetics, suggesting a possible feedback mechanism for regulating enzyme activity. Structural comparisons between members of this enzyme class give deeper insights into conserved features important for catalysis.
Organizational Affiliation:
Structural Genomics Consortium, Botnar Research Centre, University of Oxford, Oxford OX3 7LD, United Kingdom. Electronic address: kate.kavanagh@sgc.ox.ac.uk.