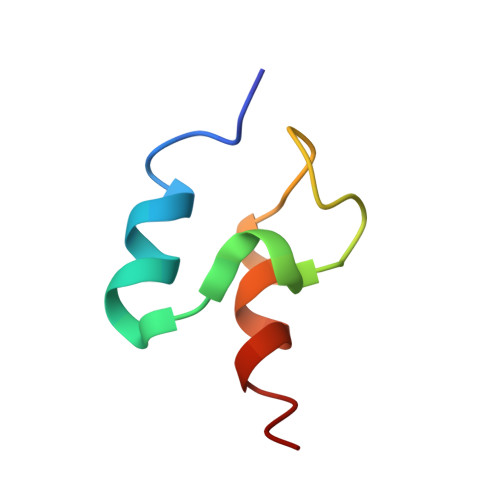
The high-resolution structure of the peripheral subunit-binding domain of dihydrolipoamide acetyltransferase from the pyruvate dehydrogenase multienzyme complex of Bacillus stearothermophilus.
Kalia, Y.N., Brocklehurst, S.M., Hipps, D.S., Appella, E., Sakaguchi, K., Perham, R.N.(1993) J Mol Biol 230: 323-341
- PubMed: 8450544
- DOI: https://doi.org/10.1006/jmbi.1993.1145
- Primary Citation of Related Structures:
2PDD, 2PDE - PubMed Abstract:
The three-dimensional structure of a 43-residue active, synthetic peptide encompassing the peripheral subunit-binding domain of dihydrolipoamide acetyltransferase from the pyruvate dehydrogenase multienzyme complex of Bacillus stearothermophilus has been determined by means of a multi-cooling dynamical simulated annealing protocol using restraints derived from 1H nuclear magnetic resonance spectroscopy. A total of 442 experimentally derived restraints including 13 dihedral angle (phi, chi 1) restraints were used. A final set of 35 structures was calculated with a root-mean-square deviation from the mean co-ordinates of 0.36 A for the backbone atoms and 0.96 A when side-chain heavy atoms were included for the well-defined region comprising residues Val7 to Leu39. Although assignments were made and sequential connectivities observed for the N-terminal six and C-terminal four residues, the absence of long-range NOEs suggests that the terminal regions are largely unstructured. The binding domain contains two short parallel alpha-helices (residues Val7 to Lys14 and Lys32 to Leu39), a3(10)-helix (residues Asp17 to Val21) and a structured loop made up of overlapping beta-turns (residues Gln22 to Leu31), which enclose a close-packed hydrophobic core. The loop is stabilized to a large extent by Asp34. This residue is conserved in all peripheral subunit-binding domains and its carboxylate side-chain forms a set of side-chain-main-chain hydrogen bonds with the main-chain amide protons of Gly23, Thr24, Gly25 and Leu31 and a side-chain-side-chain hydrogen bond with the hydroxyl group of Thr24. We propose that a peripheral subunit-binding site may be located in the loop region, which contains a series of highly conserved residues and provides a number of potential recognition sites. The structured region of the binding domain, comprising 33 residues, represents an exceptionally short amino acid sequence with defined tertiary structure that has no disulphide bond, ligand or cofactor to stabilize the fold. It may be approaching the lower size limit for a three-dimensional structure possessing features characteristic of larger structures, including a close-packed, non-polar interior. The organization of the side-chains in the hydrophobic core may have implications for de novo protein design.
Organizational Affiliation:
Department of Biochemistry, University of Cambridge, U.K.