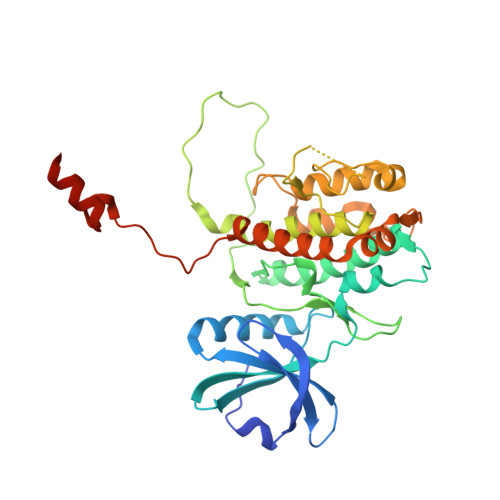
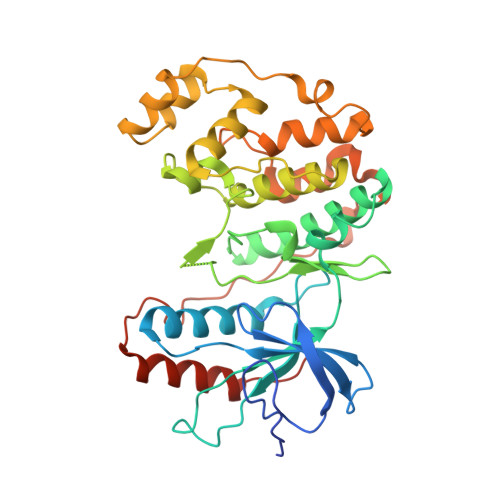
Molecular basis of MAPK-activated protein kinase 2:p38 assembly
White, A., Pargellis, C.A., Studts, J.M., Werneburg, B.G., Farmer II, B.T.(2007) Proc Natl Acad Sci U S A 104: 6353-6358
- PubMed: 17395714
- DOI: https://doi.org/10.1073/pnas.0701679104
- Primary Citation of Related Structures:
2OZA - PubMed Abstract:
p38 MAPK and MAPK-activated protein kinase 2 (MK2) are key components of signaling pathways leading to many cellular responses, notably the proinflammatory cytokine production. The physical association of p38alpha isoform and MK2 is believed to be physiologically important for this signaling. We report the 2.7-A resolution crystal structure of the unphosphorylated complex between p38alpha and MK2. These protein kinases bind "head-to-head," present their respective active sites on approximately the same side of the heterodimer, and form extensive intermolecular interactions. Among these interactions, the MK2 Ile-366-Ala-390, which includes the bipartite nuclear localization signal, binds to the p38alpha-docking region. This binding supports the involvement of noncatalytic regions to the tight binding of the MK2:p38alpha binary assembly. The MK2 residues 345-365, containing the nuclear export signal, block access to the p38alpha active site. Some regulatory phosphorylation regions of both protein kinases engage in multiple interactions with one another in this complex. This structure gives new insights into the regulation of the protein kinases p38alpha and MK2, aids in the better understanding of their known cellular and biochemical studies, and provides a basis for understanding other regulatory protein-protein interactions involving signal transduction proteins.
Organizational Affiliation:
Department of Medicinal Chemistry, Boehringer Ingelheim Pharmaceuticals, Inc., Ridgefield, CT 06877, USA. awhite@rdg.hoehringer-ingelheim.com