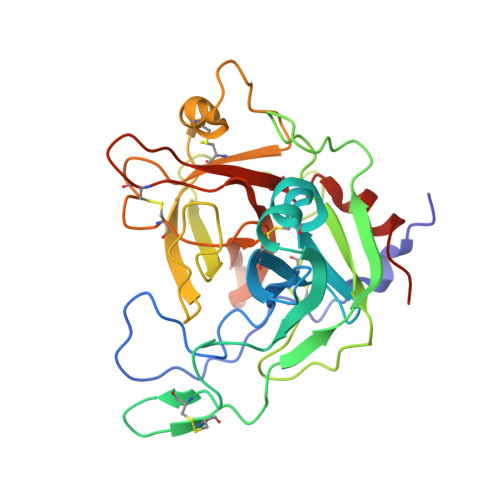
Crystal structure of the serine protease domain of prophenoloxidase activating factor-I
Piao, S., Kim, S., Kim, J.H., Park, J.W., Lee, B.L., Ha, N.C.(2007) J Biol Chem 282: 10783-10791
- PubMed: 17287215
- DOI: https://doi.org/10.1074/jbc.M611556200
- Primary Citation of Related Structures:
2OLG - PubMed Abstract:
A family of serine proteases (SPs) mediates the proteolytic cascades of embryonic development and immune response in invertebrates. These proteases, called easter-type SPs, consist of clip and chymotrypsin-like SP domains. The SP domain of easter-type proteases differs from those of typical SPs in its primary structure. Herein, we report the first crystal structure of the SP domain of easter-type proteases, presented as that of prophenoloxidase activating factor (PPAF)-I in zymogen form. This structure reveals several important structural features including a bound calcium ion, an additional loop with a unique disulfide linkage, a canyon-like deep active site, and an exposed activation loop. We subsequently show the role of the bound calcium and the proteolytic susceptibility of the activation loop, which occurs in a clip domain-independent manner. Based on biochemical study in the presence of heparin, we suggest that PPAF-III, highly homologous to PPAF-I, contains a surface patch that is responsible for enhancing the catalytic activity through interaction with a nonsubstrate region of a target protein. These results provide insights into an activation mechanism of easter-type proteases in proteolytic cascades, in comparison with the well studied blood coagulation enzymes in mammals.
Organizational Affiliation:
College of Pharmacy and Research Institute for Drug Development, Pusan National University, Jangjeon-dong, Geumjeong-gu, Busan 609-735, Korea.