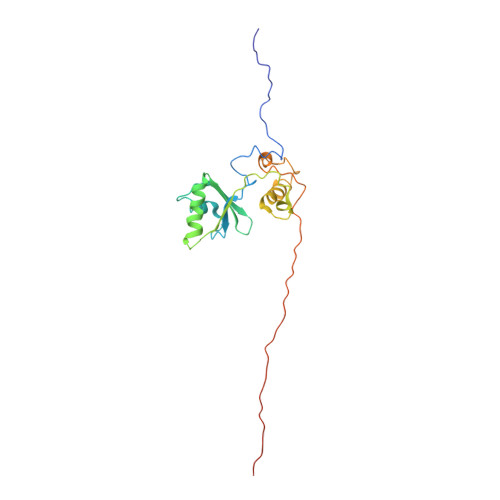
Structure and semi-sequence-specific RNA binding of Nrd1.
Bacikova, V., Pasulka, J., Kubicek, K., Stefl, R.(2014) Nucleic Acids Res 42: 8024-8038
- PubMed: 24860164
- DOI: https://doi.org/10.1093/nar/gku446
- Primary Citation of Related Structures:
2M88 - PubMed Abstract:
In Saccharomyces cerevisiae, the Nrd1-dependent termination and processing pathways play an important role in surveillance and processing of non-coding ribonucleic acids (RNAs). The termination and subsequent processing is dependent on the Nrd1 complex consisting of two RNA-binding proteins Nrd1 and Nab3 and Sen1 helicase. It is established that Nrd1 and Nab3 cooperatively recognize specific termination elements within nascent RNA, GUA[A/G] and UCUU[G], respectively. Interestingly, some transcripts do not require GUA[A/G] motif for transcription termination in vivo and binding in vitro, suggesting the existence of alternative Nrd1-binding motifs. Here we studied the structure and RNA-binding properties of Nrd1 using nuclear magnetic resonance (NMR), fluorescence anisotropy and phenotypic analyses in vivo. We determined the solution structure of a two-domain RNA-binding fragment of Nrd1, formed by an RNA-recognition motif and helix-loop bundle. NMR and fluorescence data show that not only GUA[A/G] but also several other G-rich and AU-rich motifs are able to bind Nrd1 with affinity in a low micromolar range. The broad substrate specificity is achieved by adaptable interaction surfaces of the RNA-recognition motif and helix-loop bundle domains that sandwich the RNA substrates. Our findings have implication for the role of Nrd1 in termination and processing of many non-coding RNAs arising from bidirectional pervasive transcription.
Organizational Affiliation:
CEITEC-Central European Institute of Technology, Masaryk University, Brno 62500, Czech Republic National Centre for Biomolecular Research, Faculty of Science, Masaryk University, Brno 62500, Czech Republic.