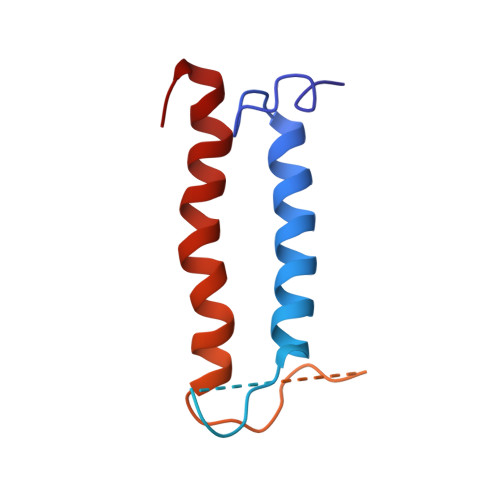
Membrane domain structures of three classes of histidine kinase receptors by cell-free expression and rapid NMR analysis.
Maslennikov, I., Klammt, C., Hwang, E., Kefala, G., Okamura, M., Esquivies, L., Mors, K., Glaubitz, C., Kwiatkowski, W., Jeon, Y.H., Choe, S.(2010) Proc Natl Acad Sci U S A 107: 10902-10907
- PubMed: 20498088
- DOI: https://doi.org/10.1073/pnas.1001656107
- Primary Citation of Related Structures:
2KSD, 2KSE, 2KSF - PubMed Abstract:
NMR structural studies of membrane proteins (MP) are hampered by complications in MP expression, technical difficulties associated with the slow process of NMR spectral peak assignment, and limited distance information obtainable for transmembrane (TM) helices. To overcome the inherent challenges in the determination of MP structures, we have developed a rapid and cost-efficient strategy that combines cell-free (CF) protein synthesis, optimized combinatorial dual-isotope labeling for nearly instant resonance assignment, and fast acquisition of long-distance information using paramagnetic probes. Here we report three backbone structures for the TM domains of the three classes of Escherichia coli histidine kinase receptors (HKRs). The ArcB and QseC TM domains are both two-helical motifs, whereas the KdpD TM domain comprises a four-helical bundle with shorter second and third helices. The interhelical distances (up to 12 A) reveal weak interactions within the TM domains of all three receptors. Determined consecutively within 8 months, these structures offer insight into the abundant and underrepresented in the Protein Data Bank class of 2-4 TM crossers and demonstrate the efficiency of our CF combinatorial dual-labeling strategy, which can be applied to solve MP structures in high numbers and at a high speed. Our results greatly expand the current knowledge of HKR structure, opening the doors to studies on their widespread and pharmaceutically important bacterial signaling mechanism.
Organizational Affiliation:
Structural Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, CA 92037, USA.