Design and structure of an equilibrium protein folding intermediate. A hint into dynamical regions of proteins
Ayuso-Tejedor, S., Angarica, V.E., Bueno, M., Campos, L.A., Abian, O., Bernado, P., Sancho, J., Jimenez, M.A.(2010) J Mol Biol 400: 922-934
- PubMed: 20553732
- DOI: https://doi.org/10.1016/j.jmb.2010.05.050
- Primary Citation of Related Structures:
2KQU - PubMed Abstract:
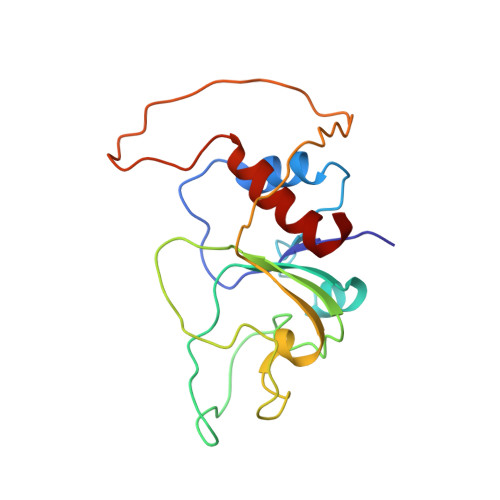
Partly unfolded protein conformations close to the native state may play important roles in protein function and in protein misfolding. Structural analyses of such conformations which are essential for their fully physicochemical understanding are complicated by their characteristic low populations at equilibrium. We stabilize here with a single mutation the equilibrium intermediate of apoflavodoxin thermal unfolding and determine its solution structure by NMR. It consists of a large native region identical with that observed in the X-ray structure of the wild-type protein plus an unfolded region. Small-angle X-ray scattering analysis indicates that the calculated ensemble of structures is consistent with the actual degree of expansion of the intermediate. The unfolded region encompasses discontinuous sequence segments that cluster in the 3D structure of the native protein forming the FMN cofactor binding loops and the binding site of a variety of partner proteins. Analysis of the apoflavodoxin inner interfaces reveals that those becoming destabilized in the intermediate are more polar than other inner interfaces of the protein. Natively folded proteins contain hydrophobic cores formed by the packing of hydrophobic surfaces, while natively unfolded proteins are rich in polar residues. The structure of the apoflavodoxin thermal intermediate suggests that the regions of natively folded proteins that are easily responsive to thermal activation may contain cores of intermediate hydrophobicity.
Organizational Affiliation:
Departamento de Bioquímica y Biología Molecular, Facultad de Ciencias, Universidad de Zaragoza, 50009 Zaragoza, Spain.