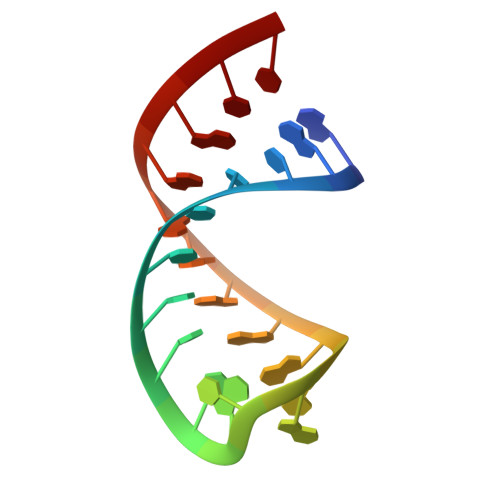
Guanidinoneomycin B Recognition of an HIV-1 RNA Helix.
Staple, D.W., Venditti, V., Niccolai, N., Elson-Schwab, L., Tor, Y., Butcher, S.E.(2008) Chembiochem 9: 93-102
- PubMed: 18058789
- DOI: https://doi.org/10.1002/cbic.200700251
- Primary Citation of Related Structures:
2JUK - PubMed Abstract:
Aminoglycoside antibiotics are small-molecule drugs that bind RNA. The affinity and specificity of aminoglycoside binding to RNA can be increased through chemical modification, such as guanidinylation. Here, we report the binding of guanidinoneomycin B (GNB) to an RNA helix from the HIV-1 frameshift site. The binding of GNB increases the melting temperature (T(m)) of the frameshift-site RNA by at least 10 degrees C, to a point at which a melting transition is not even observed in 2 M urea. A structure of the complex was obtained by using multidimensional heteronuclear NMR spectroscopic methods. We also used a novel paramagnetic-probe assay to identify the site of GNB binding to the surface of the RNA. GNB makes major-groove contacts to two sets of Watson-Crick bases and is in van der Waals contact with a highly structured ACAA tetraloop. Rings I and II of GNB fit into the major groove and form the binding interface with the RNA, whereas rings III and IV are exposed to the solvent and disordered. The binding of GNB causes a broadening of the major groove across the binding site.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin-Madison, 433 Babcock Drive, Madison, WI 53706, USA.