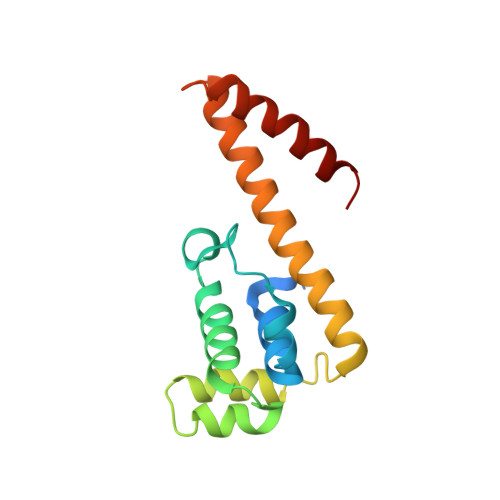
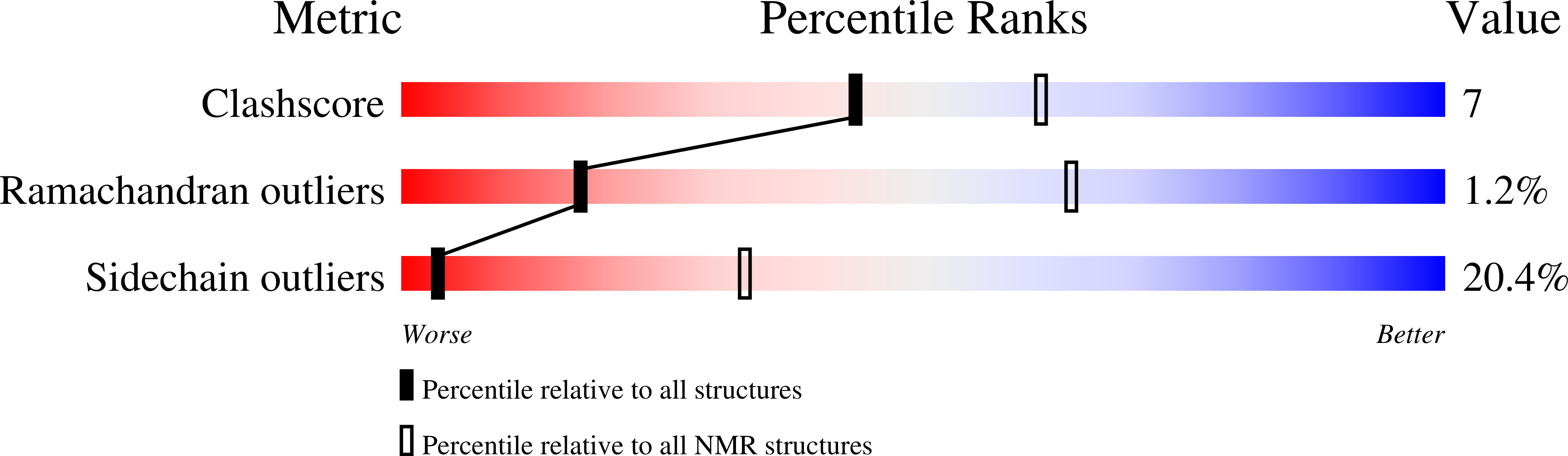Monomeric solution structure of the helicase-binding domain of Escherichia coli DnaG primase
Su, X.C., Schaeffer, P.M., Loscha, K.V., Gan, P.H.P., Dixon, N.E., Otting, G.(2006) FEBS J 273: 4997-5009
- PubMed: 17010164
- DOI: https://doi.org/10.1111/j.1742-4658.2006.05495.x
- Primary Citation of Related Structures:
2HAJ - PubMed Abstract:
DnaG is the primase that lays down RNA primers on single-stranded DNA during bacterial DNA replication. The solution structure of the DnaB-helicase-binding C-terminal domain of Escherichia coli DnaG was determined by NMR spectroscopy at near-neutral pH. The structure is a rare fold that, besides occurring in DnaG C-terminal domains, has been described only for the N-terminal domain of DnaB. The C-terminal helix hairpin present in the DnaG C-terminal domain, however, is either less stable or absent in DnaB, as evidenced by high mobility of the C-terminal 35 residues in a construct comprising residues 1-171. The present structure identifies the previous crystal structure of the E. coli DnaG C-terminal domain as a domain-swapped dimer. It is also significantly different from the NMR structure reported for the corresponding domain of DnaG from the thermophile Bacillus stearothermophilus. NMR experiments showed that the DnaG C-terminal domain does not bind to residues 1-171 of the E. coli DnaB helicase with significant affinity.
Organizational Affiliation:
Research School of Chemistry, Australian National University, Canberra, ACT 0200, Australia.