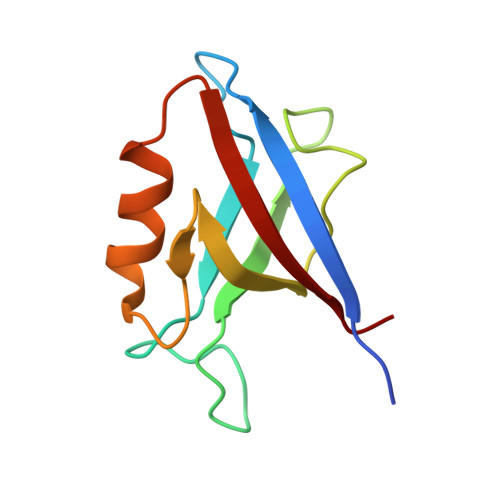
Comparative structural analysis of the Erbin PDZ domain and the first PDZ domain of ZO-1. Insights into determinants of PDZ domain specificity.
Appleton, B.A., Zhang, Y., Wu, P., Yin, J.P., Hunziker, W., Skelton, N.J., Sidhu, S.S., Wiesmann, C.(2006) J Biol Chem 281: 22312-22320
- PubMed: 16737969
- DOI: https://doi.org/10.1074/jbc.M602901200
- Primary Citation of Related Structures:
2H2B, 2H2C, 2H3L, 2H3M - PubMed Abstract:
We report a structural comparison of the first PDZ domain of ZO-1 (ZO1-PDZ1) and the PDZ domain of Erbin (Erbin-PDZ). Although the binding profile of Erbin-PDZ is extremely specific ([D/E][T/S]WV(COOH)), that of ZO1-PDZ1 is similar ([R/K/S/T][T/S][W/Y][V/I/L](COOH)) but broadened by increased promiscuity for three of the last four ligand residues. Consequently, the biological function of ZO-1 is also broadened, as it interacts with both tight and adherens junction proteins, whereas Erbin is restricted to adherens junctions. Structural analyses reveal that the differences in specificity can be accounted for by two key differences in primary sequence. A reduction in the size of the hydrophobic residue at the base of the site(0) pocket enables ZO1-PDZ1 to accommodate larger C-terminal residues. A single additional difference alters the specificity of both site(-1) and site(-3). In ZO1-PDZ1, an Asp residue makes favorable interactions with both Tyr(-1) and Lys/Arg(-3). In contrast, Erbin-PDZ contains an Arg at the equivalent position, and this side chain cannot accommodate either Tyr(-1) or Lys/Arg(-3) but, instead, interacts favorably with Glu/Asp(-3). We propose a model for ligand recognition that accounts for interactions extending across the entire binding site but that highlights several key specificity switches within the PDZ domain fold.
Organizational Affiliation:
Department of Protein Engineering, Genentech, Inc., South San Francisco, California 94080.