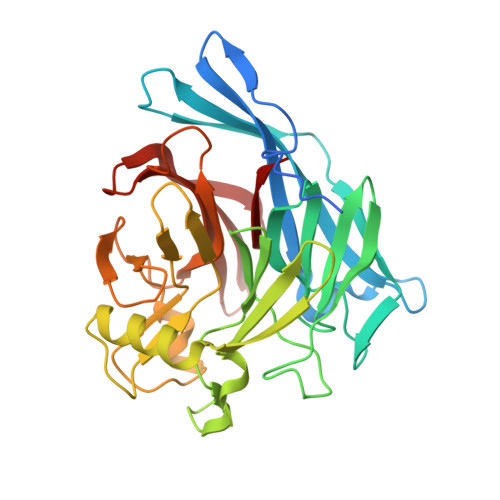
Calcium-dependent dimerization of human soluble calcium activated nucleotidase: characterization of the dimer interface.
Yang, M., Horii, K., Herr, A.B., Kirley, T.L.(2006) J Biol Chem 281: 28307-28317
- PubMed: 16835225
- DOI: https://doi.org/10.1074/jbc.M604413200
- Primary Citation of Related Structures:
2H2N, 2H2U - PubMed Abstract:
Mammals express a protein homologous to soluble nucleotidases used by blood-sucking insects to inhibit host blood clotting. These vertebrate nucleotidases may play a role in protein glycosylation. The activity of this enzyme family is strictly dependent on calcium, which induces a conformational change in the secreted, soluble human nucleotidase. The crystal structure of this human enzyme was recently solved; however, the mechanism of calcium activation and the basis for the calcium-induced changes remain unclear. In this study, using analytical ultracentrifugation and chemical cross-linking, we show that calcium or strontium induce noncovalent dimerization of the soluble human enzyme. The location and nature of the dimer interface was elucidated using a combination of site-directed mutagenesis and chemical cross-linking, coupled with crystallographic analyses. Replacement of Ile(170), Ser(172), and Ser(226) with cysteine residues resulted in calcium-dependent, sulfhydryl-specific intermolecular cross-linking, which was not observed after cysteine introduction at other surface locations. Analysis of a super-active mutant, E130Y, revealed that this mutant dimerized more readily than the wild-type enzyme. The crystal structure of the E130Y mutant revealed that the mutated residue is found in the dimer interface. In addition, expression of the full-length nucleotidase revealed that this membrane-bound form can also dimerize and that these dimers are stabilized by spontaneous oxidative cross-linking of Cys(30), located between the single transmembrane helix and the start of the soluble sequence. Thus, calcium-mediated dimerization may also represent a mechanism for regulation of the activity of this nucleotidase in the physiological setting of the endoplasmic reticulum or Golgi.
Organizational Affiliation:
Department of Pharmacology and Cell Biophysics, College of Medicine, University of Cincinnati College of Medicine, Ohio 45267-0575, USA.