Molecular basis of inhibition of the ribonuclease activity in colicin e5 by its cognate immunity protein
Luna-Chavez, C., Lin, Y.L., Huang, R.H.(2006) J Mol Biol 358: 571-579
- PubMed: 16524591
- DOI: https://doi.org/10.1016/j.jmb.2006.02.014
- Primary Citation of Related Structures:
2FHZ - PubMed Abstract:
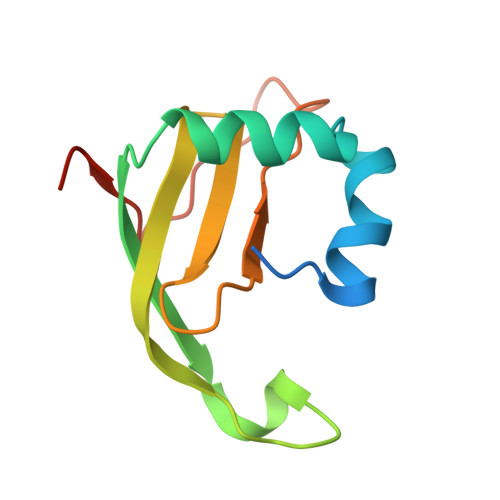
Colicin E5 is a tRNA-specific ribonuclease that recognizes and cleaves four tRNAs in Escherichia coli that contain the hypermodified nucleoside queuosine (Q) at the wobble position. Cells that produce colicin E5 also synthesize the cognate immunity protein (Im5) that rapidly and tightly associates with colicin E5 to prevent it from cleaving its own tRNAs to avoid suicide. We report here the crystal structure of Im5 in a complex with the activity domain of colicin E5 (E5-CRD) at 1.15A resolution. The structure reveals an extruded domain from Im5 that docks into the recessed RNA binding cleft in E5-CRD, resulting in extensive interactions between the two proteins. The interactions are primarily hydrophilic, with an interface that contains complementary surface charges between the two proteins. Detailed interactions in three separate regions of the interface account for specific recognition of colicin E5 by Im5. Furthermore, single-site mutational studies of Im5 confirmed the important role of particular residues in recognition and binding of colicin E5. Structural comparison of the complex reported here with E5-CRD alone, as well as with a docking model of RNA-E5-CRD, indicates that Im5 achieves its inhibition by physically blocking the cleft in colicin E5 that engages the RNA substrate.
Organizational Affiliation:
Center for Biophysics and Computational Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA.