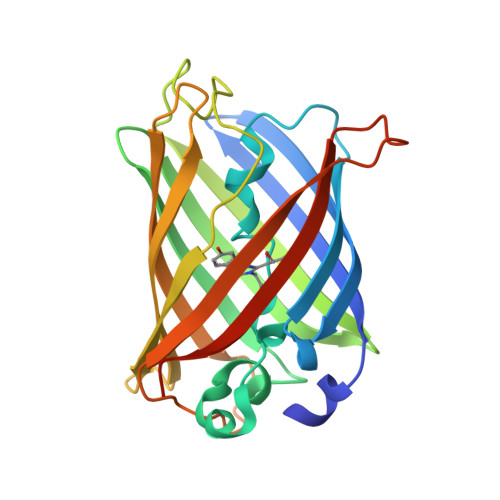
Ultrafast excited-state dynamics in the green fluorescent protein variant S65T/H148D. 1. Mutagenesis and structural studies.
Shu, X., Kallio, K., Shi, X., Abbyad, P., Kanchanawong, P., Childs, W., Boxer, S.G., Remington, S.J.(2007) Biochemistry 46: 12005-12013
- PubMed: 17918959
- DOI: https://doi.org/10.1021/bi7009037
- Primary Citation of Related Structures:
2DUE, 2DUF, 2DUG, 2DUH, 2DUI - PubMed Abstract:
Wild type green fluorescent protein (wt-GFP) and the variant S65T/H148D each exhibit two absorption bands, A and B, which are associated with the protonated and deprotonated chromophores, respectively. Excitation of either band leads to green emission. In wt-GFP, excitation of band A ( approximately 395 nm) leads to green emission with a rise time of 10-15 ps, due to excited-state proton transfer (ESPT) from the chromophore hydroxyl group to an acceptor. This process produces an anionic excited-state intermediate I* that subsequently emits a green photon. In the variant S65T/H148D, the A band absorbance maximum is red-shifted to approximately 415 nm, and as detailed in the accompanying papers, when the A band is excited, green fluorescence appears with a rise time shorter than the instrument time resolution ( approximately 170 fs). On the basis of the steady-state spectroscopy and high-resolution crystal structures of several variants described herein, it is proposed that in S65T/H148D, the red shift of absorption band A and the ultrafast appearance of green fluorescence upon excitation of band A are due to a very short (
Organizational Affiliation:
Institute of Molecular Biology and Department of Physics, University of Oregon, Eugene, Oregon 97403-1229, USA.