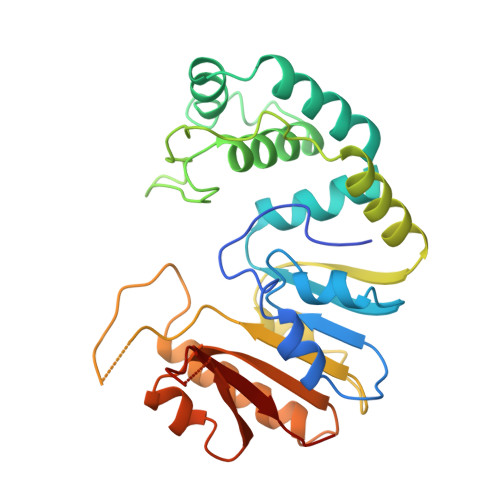
Crystal structure of the DpnM DNA adenine methyltransferase from the DpnII restriction system of streptococcus pneumoniae bound to S-adenosylmethionine.
Tran, P.H., Korszun, Z.R., Cerritelli, S., Springhorn, S.S., Lacks, S.A.(1998) Structure 6: 1563-1575
- PubMed: 9862809
- DOI: https://doi.org/10.1016/s0969-2126(98)00154-3
- Primary Citation of Related Structures:
2DPM - PubMed Abstract:
. Methyltransferases (Mtases) catalyze the transfer of methyl groups from S-adenosylmethionine (AdoMet) to a variety of small molecular and macromolecular substrates. These enzymes contain a characteristic alpha/beta structural fold. Four groups of DNA Mtases have been defined and representative structures have been determined for three groups. DpnM is a DNA Mtase that acts on adenine N6 in the sequence GATC; the enzyme represents group alpha DNA Mtases, for which no structures are known. . The structure of DpnM in complex with AdoMet was determined at 1.80 A resolution. The protein comprises a consensus Mtase fold with a helical cluster insert. DpnM binds AdoMet in a similar manner to most other Mtases and the enzyme contains a hollow that can accommodate DNA. The helical cluster supports a shelf within the hollow that may recognize the target sequence. Modeling studies indicate a potential site for binding the target adenine, everted from the DNA helix. Comparison of the DpnM structure and sequences of group alpha DNA Mtases indicates that the group is a genetically related family. Structural comparisons show DpnM to be most similar to a small-molecule Mtase and then to macromolecular Mtases, although several dehydrogenases show greater similarity than one DNA Mtase. . DpnM, and by extension the DpnM family or group alpha Mtases, contains the consensus fold and AdoMet-binding motifs found in most Mtases. Structural considerations suggest that macromolecular Mtases evolved from small-molecule Mtases, with different groups of DNA Mtases evolving independently. Mtases may have evolved from dehydrogenases. Comparison of these enzymes indicates that in protein evolution, the structural fold is most highly conserved, then function and lastly sequence.
Organizational Affiliation:
Department of Biology, Brookhaven National Laboratory, Upton, NY 11973,USA.