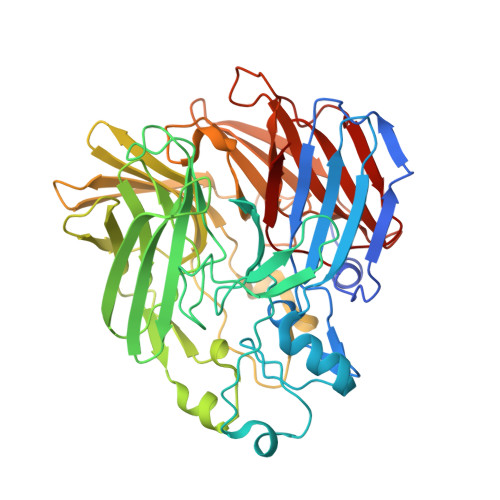
The Structure of a Retinal-Forming Carotenoid Oxygenase
Kloer, D.P., Ruch, S., Al-Babili, S., Beyer, P., Schulz, G.E.(2005) Science 308: 267
- PubMed: 15821095
- DOI: https://doi.org/10.1126/science.1108965
- Primary Citation of Related Structures:
2BIW, 2BIX - PubMed Abstract:
Enzymes that produce retinal and related apocarotenoids constitute a sequence- and thus structure-related family, a member of which was analyzed by x-ray diffraction. This member is an oxygenase and contains an Fe2+-4-His arrangement at the axis of a seven-bladed beta-propeller chain fold covered by a dome formed by six large loops. The Fe2+ is accessible through a long nonpolar tunnel that holds a carotenoid derivative in one of the crystals. On binding, three consecutive double bonds of this carotenoid changed from a straight all-trans to a cranked cis-trans-cis conformation. The remaining trans bond is located at the dioxygen-ligated Fe2+ and cleaved by oxygen.
Organizational Affiliation:
Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Albertstrasse 21, 79104 Freiburg im Breisgau, Germany.