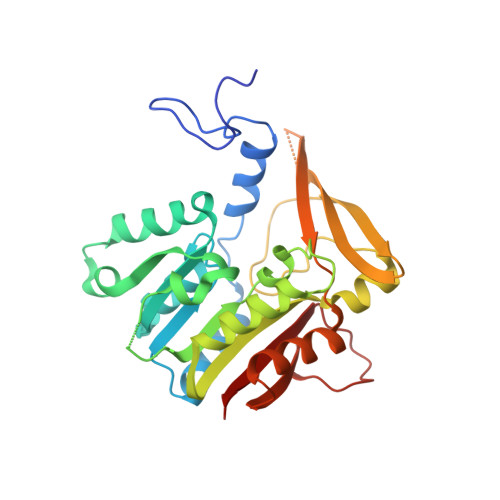
Destabilization of human glycine N-methyltransferase by H176N mutation.
Luka, Z., Pakhomova, S., Luka, Y., Newcomer, M.E., Wagner, C.(2007) Protein Sci 16: 1957-1964
- PubMed: 17660255
- DOI: https://doi.org/10.1110/ps.072921507
- Primary Citation of Related Structures:
2AZT - PubMed Abstract:
In the presence of moderate (2-4 M) urea concentrations the tetrameric enzyme, glycine N-methyltransferase (GNMT), dissociates into compact monomers. Higher concentrations of urea (7-8 M) promote complete denaturation of the enzyme. We report here that the H176N mutation in this enzyme, found in humans with hypermethioninaemia, significantly decreases stability of the tetramer, although H176 is located far from the intersubunit contact areas. Dissociation of the tetramer to compact monomers and unfolding of compact monomers of the mutant protein were detected by circular dichroism, quenching of fluorescence emission, size-exclusion chromatography, and enzyme activity. The values of apparent free energy of dissociation of tetramer and of unfolding of compact monomers for the H176N mutant (27.7 and 4.2 kcal/mol, respectively) are lower than those of wild-type protein (37.5 and 6.2 kcal/mol). A 2.7 A resolution structure of the mutant protein revealed no significant difference in the conformation of the protein near the mutated residue.
Organizational Affiliation:
Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, Tennessee 37232, USA.