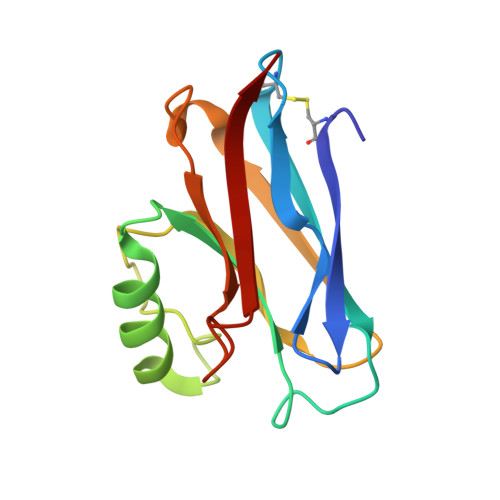
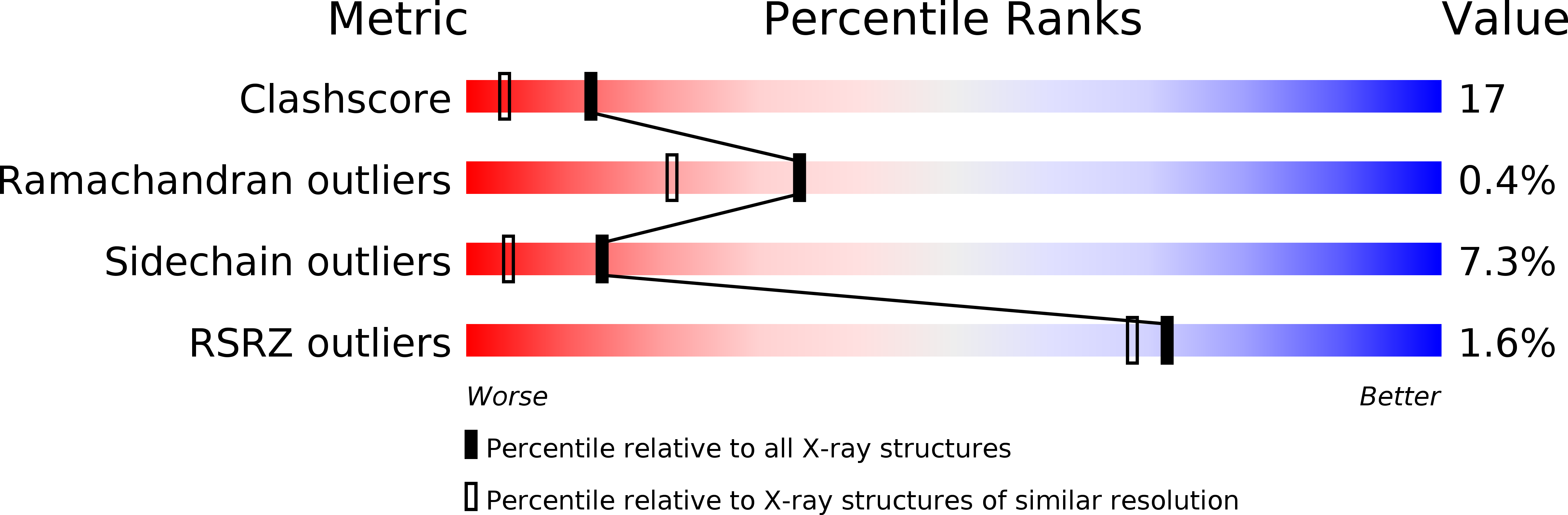Structure of azurin from Alcaligenes denitrificans refinement at 1.8 A resolution and comparison of the two crystallographically independent molecules.
Baker, E.N.(1988) J Mol Biol 203: 1071-1095
- PubMed: 3210236
- DOI: https://doi.org/10.1016/0022-2836(88)90129-5
- Primary Citation of Related Structures:
2AZA - PubMed Abstract:
The structure of the blue copper protein azurin, from Alcaligenes denitrificans, has been refined crystallographically by restrained least-squares methods. The final crystallographic R value for 21,980 observed reflections to 1.8 A (1 A = 0.1 nm) resolution is 0.157. The asymmetric unit of the crystal contains two independent azurin molecules, the model for which comprises 1973 protein atoms, together with three SO2-4 ions, and 281 water molecules. Comparison of the two molecules shows very high correspondence. For 125 out of 129 residues (excluding only the chain termini, residues 1 to 2 and 128 to 129) the root-mean-square (r.m.s.) deviation in main-chain atom positions is 0.27 A. For other structural parameters r.m.s. deviations are also low; torsion angles 6.5 degrees, hydrogen bond lengths 0.12 A, bonds to copper 0.04 A and bond angles at the copper 3.9 degrees. The only significant differences are at the chain termini and in several loops. Some of these can be attributed to crystal packing effects, others to genuine structural microheterogeneity. Refinement has confirmed that the copper co-ordination is best described as distorted trigonal planar, with strong in-plane bonds to His46 N delta 1, His117 N delta 1 and Cys112 S gamma, and much weaker axial interactions with Met121 S delta and Gly45 C = O. Two N-H...S hydrogen bonds characterize Cys112 S gamma as a thiolate (S-) sulphur and may influence the visible absorption maximum. Atoms in and around the copper site have very low mobility, whereas the most mobile regions of the molecule are the chain termini and some of the connecting loops between secondary structure elements, especially those at the "southern" end, remote from the copper site. Main-chain to side-chain hydrogen bonds supply important stabilizing interactions at the "northern" end. Surface features include the hydrophobic patch around His117, probably important for electron transfer, the SO2-4 site at His83, and the general absence of ion pairs, despite the presence of many charged amino acid residues. The 281 water molecules include 182 that occur as approximately twofold-related pairs. There are no internal water molecules. The water sites common to both azurin molecules include those in surface pockets and some in intermolecular contact regions. They are characterized by relatively low thermal parameters and numerous protein contacts.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Massey University, Palmerston North, New Zealand.