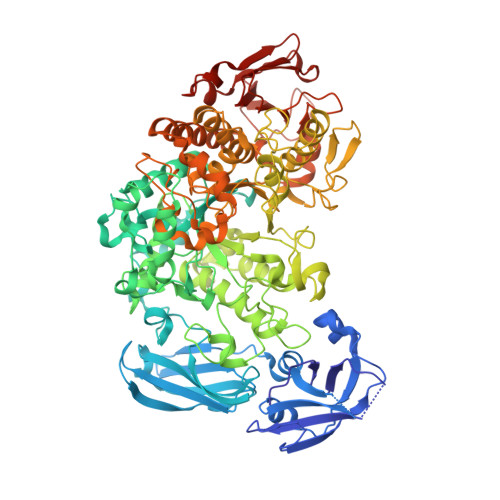
Crystal Structure of an Essential Enzyme in Seed Starch Degradation: Barley Limit Dextrinase in Complex with Cyclodextrins.
Vester-Christensen, M.B., Abou Hachem, M., Svensson, B., Henriksen, A.(2010) J Mol Biol 403: 739
- PubMed: 20863834
- DOI: https://doi.org/10.1016/j.jmb.2010.09.031
- Primary Citation of Related Structures:
2Y4S, 2Y5E - PubMed Abstract:
Barley limit dextrinase [Hordeum vulgare limit dextrinase (HvLD)] catalyzes the hydrolysis of α-1,6 glucosidic linkages in limit dextrins. This activity plays a role in starch degradation during germination and presumably in starch biosynthesis during grain filling. The crystal structures of HvLD in complex with the competitive inhibitors α-cyclodextrin (CD) and β-CD are solved and refined to 2.5 Å and 2.1 Å, respectively, and are the first structures of a limit dextrinase. HvLD belongs to glycoside hydrolase 13 family and is composed of four domains: an immunoglobulin-like N-terminal eight-stranded β-sandwich domain, a six-stranded β-sandwich domain belonging to the carbohydrate binding module 48 family, a catalytic (β/α)(8)-like barrel domain that lacks α-helix 5, and a C-terminal eight-stranded β-sandwich domain of unknown function. The CDs are bound at the active site occupying carbohydrate binding subsites +1 and +2. A glycerol and three water molecules mimic a glucose residue at subsite -1, thereby identifying residues involved in catalysis. The bulky Met440, a unique residue at its position among α-1,6 acting enzymes, obstructs subsite -4. The steric hindrance observed is proposed to affect substrate specificity and to cause a low activity of HvLD towards amylopectin. An extended loop (Asp513-Asn520) between β5 and β6 of the catalytic domain also seems to influence substrate specificity and to give HvLD a higher affinity for α-CD than pullulanases. The crystal structures additionally provide new insight into cation sites and the concerted action of the battery of hydrolytic enzymes in starch degradation.
Organizational Affiliation:
Enzyme and Protein Chemistry, Department of Systems Biology, Søltofts Plads, Building 224, Technical University of Denmark, Kgs. Lyngby, Denmark.