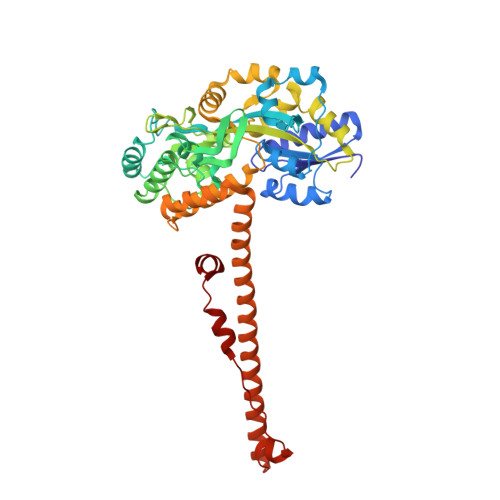
Charge-Surrounded Pockets and Electrostatic Interactions with Small Ions Modulate the Activity of Retroviral Fusion Proteins.
Lamb, D., Schuttelkopf, A.W., Van Aalten, D.M.F., Brighty, D.W.(2011) PLoS Pathog 7: 1268
- PubMed: 21304939
- DOI: https://doi.org/10.1371/journal.ppat.1001268
- Primary Citation of Related Structures:
2XZ3 - PubMed Abstract:
Refolding of viral class-1 membrane fusion proteins from a native state to a trimer-of-hairpins structure promotes entry of viruses into cells. Here we present the structure of the bovine leukaemia virus transmembrane glycoprotein (TM) and identify a group of asparagine residues at the membrane-distal end of the trimer-of-hairpins that is strikingly conserved among divergent viruses. These asparagines are not essential for surface display of pre-fusogenic envelope. Instead, substitution of these residues dramatically disrupts membrane fusion. Our data indicate that, through electrostatic interactions with a chloride ion, the asparagine residues promote assembly and profoundly stabilize the fusion-active structures that are required for viral envelope-mediated membrane fusion. Moreover, the BLV TM structure also reveals a charge-surrounded hydrophobic pocket on the central coiled coil and interactions with basic residues that cluster around this pocket are critical to membrane fusion and form a target for peptide inhibitors of envelope function. Charge-surrounded pockets and electrostatic interactions with small ions are common among class-1 fusion proteins, suggesting that small molecules that specifically target such motifs should prevent assembly of the trimer-of-hairpins and be of value as therapeutic inhibitors of viral entry.
Organizational Affiliation:
The Biomedical Research Institute, College of Medicine, Ninewells Hospital, The University of Dundee, Dundee, UK.