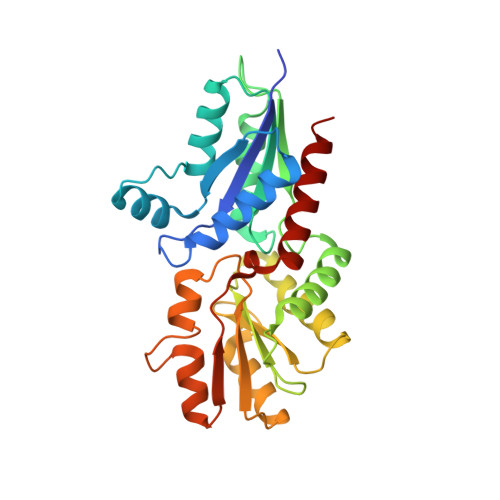
Evolution in a Family of Chelatases Facilitated by the Introduction of Active Site Asymmetry and Protein Oligomerization.
Romao, C.V., Ladakis, D., Lobo, S.A.L., Carrondo, M.A., Brindley, A.A., Deery, E., Matias, P.M., Pickersgill, R.W., Saraiva, L.M., Warren, M.J.(2011) Proc Natl Acad Sci U S A 108: 97
- PubMed: 21173279
- DOI: https://doi.org/10.1073/pnas.1014298108
- Primary Citation of Related Structures:
2XVX, 2XVZ, 2XWP, 2XWQ, 2XWS - PubMed Abstract:
The class II chelatases associated with heme, siroheme, and cobalamin biosynthesis are structurally related enzymes that insert a specific metal ion (Fe(2+) or Co(2+)) into the center of a modified tetrapyrrole (protoporphyrin or sirohydrochlorin). The structures of two related class II enzymes, CbiX(S) from Archaeoglobus fulgidus and CbiK from Salmonella enterica, that are responsible for the insertion of cobalt along the cobalamin biosynthesis pathway are presented in complex with their metallated product. A further structure of a CbiK from Desulfovibrio vulgaris Hildenborough reveals how cobalt is bound at the active site. The crystal structures show that the binding of sirohydrochlorin is distinctly different to porphyrin binding in the protoporphyrin ferrochelatases and provide a molecular overview of the mechanism of chelation. The structures also give insights into the evolution of chelatase form and function. Finally, the structure of a periplasmic form of Desulfovibrio vulgaris Hildenborough CbiK reveals a novel tetrameric arrangement of its subunits that are stabilized by the presence of a heme b cofactor. Whereas retaining colbaltochelatase activity, this protein has acquired a central cavity with the potential to chaperone or transport metals across the periplasmic space, thereby evolving a new use for an ancient protein subunit.
Organizational Affiliation:
Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Avenida da República (EAN), 2780-157 Oeiras, Portugal. cmromao@itqb.unl.pt