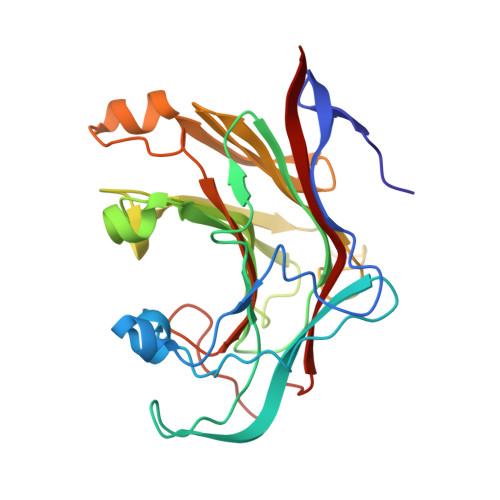
Crystal Structure of a Family 16 Endoglucanase from the Hyperthermophile Pyrococcus Furiosus-Structural Basis of Substrate Recognition.
Ilari, A., Fiorillo, A., Angelaccio, S., Florio, R., Chiaraluce, R., Van Der Oost, J., Consalvi, V.(2009) FEBS J 276: 1048
- PubMed: 19154353
- DOI: https://doi.org/10.1111/j.1742-4658.2008.06848.x
- Primary Citation of Related Structures:
2VY0 - PubMed Abstract:
Bacterial and archaeal endo-beta-1,3-glucanases that belong to glycoside hydrolase family 16 share a beta-jelly-roll fold, but differ significantly in sequence and in substrate specificity. The crystal structure of the laminarinase (EC 3.2.1.39) from the hyperthermophilic archaeon Pyrococcus furiosus (pfLamA) has been determined at 2.1 A resolution by molecular replacement. The pfLamA structure reveals a kink of six residues (72-77) at the entrance of the catalytic cleft. This peptide is absent in the endoglucanases from alkaliphilic Nocardiopsis sp. strain F96 and Bacillus macerans, two proteins displaying an overall fold similar to that of pfLamA, but with different substrate specificity. A deletion mutant of pfLamA, lacking residues 72-75, hydrolyses the mixed-linkage beta-1,3-1,4-glucan lichenan 10 times more efficiently than the wild-type protein, indicating the importance of the kink in substrate preference.
Organizational Affiliation:
CNR Institute of Molecular Biology and Pathology, Italy. andrea.ilari@uniroma1.it